General principles of cellular communication
Use 'Print preview' to check the number of pages and printer settings.
Print functionality varies between browsers.
Printable page generated Friday, 15 May 2026, 10:11 AM
General principles of cellular communication
Introduction
All organisms, whether unicellular or multicellular, need to respond to their ever-changing environment in order to survive and flourish. Such responses are governed by the ability of cells to sense physical changes and chemical cues occurring around them. The process of sensing and responding to extrinsic signals is often termed cellular communication, although scientists also use terms such as ‘signal transduction’ or ‘signalling’.
Cells respond to a wide range of extrinsic signals that include chemical messengers (e.g. hormones, growth factors, neurotransmitters), electrical impulses, mechanical forces, pH, heat and light.
In this free course, General principles of cellular communication, you will explore the most common paradigm for cellular communication, which is the detection of extrinsic stimuli by receptors on the surface of cells. Particular emphasis is placed on how the interaction between an extrinsic stimulus and its receptor on the cell surface subsequently causes cellular responses through the activation of specific intracellular signalling pathways. You will explore this chain of events using well-characterised examples in prokaryotic and eukaryotic cells.
This OpenLearn course is an adapted extract from the Open University course S317 Biological science: from genes to species.
Learning outcomes
After studying this course, you should be able to:
describe in general terms how extrinsic signals activate cell signalling pathways to elicit cellular responses
describe how cell signalling processes facilitate quorum sensing in bacteria
outline the diverse roles played by proteins in cellular communication and how various proteins can function together in a signalling pathway
relate the activities and properties (including domain structure) of proteins to their roles in signalling.
1 Diversity and evolution of cell signalling pathways
Cellular communication encompasses a vast range of extrinsic signals, intracellular signalling pathways and cellular responses. In fact, no two cell types express exactly the same repertoire of signalling components. Rather, cells have signalling systems that suit their physiological function. The main focus of this topic is cellular communication that occurs when extrinsic stimuli bind to receptors on their target cells. Although not all cellular communication relies on the activation of receptors, it is the most common mechanism by which cells sense their environment or communicate with each other. The central signalling paradigm that will be explored in this course is depicted in Figure 1.
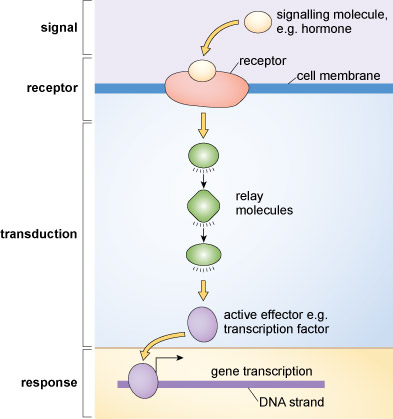
By activating receptors, extrinsic signals trigger events that relay information within cells and ultimately cause cells to change their behaviour. It is important to remember that receptors are highly selective for their specific (cognate) extrinsic stimuli. In most cases, a particular kind of receptor will only be activated by one type of extrinsic stimulus.
Activated receptors often have pleiotropic actions. That is, they alter the activity of numerous cellular processes simultaneously. These processes could include DNA transcription, protein synthesis or changes in metabolic activity. The overall effect of switching various processes on or off determines the consequent change in cellular behaviour. The fidelity, accuracy and appropriateness of these cellular communication processes are critically important for the cell and for the organism. It is well known that aberrant cellular communication leads to conditions such as cancer, diabetes, heart failure and neurological diseases.
Cells are simultaneously bombarded with numerous extrinsic signals and they make sense of these incoming signals through the activation of specific signalling pathways.
Just a few of the many signalling systems expressed in a typical mammalian cell are depicted in Figure 2. The key point that you should take from this illustration is that cellular communication triggers specific responses by recruiting particular signalling pathways. The activation of receptors following ligand binding is conveyed into a cell by a cascade of signalling proteins or messengers. Note that each pathway can elicit a variety of responses, although only one example is shown for each of the pathways in Figure 2
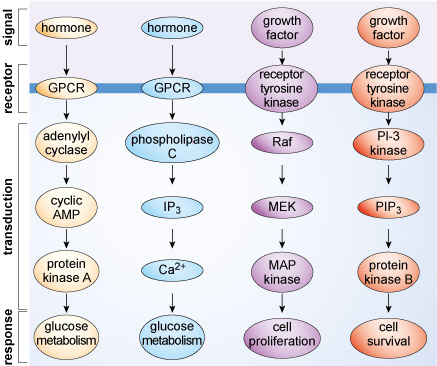
Exactly how a cell will respond to extrinsic stimuli is sometimes hard to predict, even for scientists with considerable experience of studying cellular communication. In part, the response of a cell is determined by the input signals it receives, but there are also many intrinsic factors that determine how cells respond. For example, the age of a cell, its position within the cell cycle and its metabolic status could impact on how it responds to particular extrinsic stimuli. Under some conditions, such as in a nutrient-rich environment, a cell could receive an extrinsic signal that activates anabolic processes and cell division. At another time, when nutrients are depleted, the same signal may trigger catabolism and cell death.
Although cellular signalling pathways are numerous and complex, there are in fact relatively few pathways in comparison to the diversity of cell types and their intricate molecular processes. For example, development in multicellular animals essentially relies on only seven distinct signalling pathways, commonly called hedgehog, wingless-related, transforming growth factor-β, receptor tyrosine kinases, Notch, JAK/STAT and nuclear hormone receptors (names that variously identify a key component, signal, receptor type or function). These seven pathways are used during development to define the size, shape and other characteristics of an animal. How can so few pathways achieve so much? The answer is that signalling pathways can act together to produce outcomes that are different from the outcome of single pathways acting alone. This combinatorial action of pathways may actually allow hundreds of different signalling–response combinations.
In general terms, the number of genes involved in signalling increases with complexity of an organism, with vertebrates (e.g. humans) having many more such genes than invertebrate species (Table 1). Whole genome duplication events, such as those that occurred in the evolution of vertebrates between 550 and 450 million years ago, were responsible for these marked differences. However, the correlation between complexity and number of genes involved in signalling is not a strict one. Thus, while a human generally has many more genes encoding components of the seven developmental signalling pathways than does the nematode worm Caenorhabditis elegans, the latter has many more putative nuclear hormone receptors (Table 1).
Not only have genes encoding signalling components been acquired during evolution, but such genes have also been lost. For example, the nematode C. elegans does not possess genes for the hedgehog signalling pathway that are found in other organisms (insects and other nematodes) that share a close common ancestor.
| H. sapiens (human) | D. melanogaster (fruitfly) | C. elegans (nematode worm) | S. cerevisiae (yeast) | |
|---|---|---|---|---|
| Ligands | ||||
| RTK ligand | 48 | 3 | 4 | 0 |
| TGF-β | 29 | 6 | 4 | 0 |
| Wnt | 18 | 7 | 5 | 0 |
| Notch ligand | 3 | 2 | 2 | 0 |
| Receptor | ||||
| RTK | 25 | 6 | 1 | 0 |
| Wnt receptor | 12 | 6 | 5 | 0 |
| NHR | 59 | 25 | 270 | 1 |
Footnotes
Key: RTK, receptor tyrosine kinase; TGF-β, transforming growth factor-β; Wnt, wingless-related; NHR, nuclear hormone receptorFor many of the cellular communication mechanisms found in animals there are analogous systems in plants, fungi and protists. An example is the mitogen-activated protein kinase (MAP kinase or MAPK) cascade. It is likely that ancestral eukaryotic cells developed complex signalling systems well over two billion years ago, long before multicellular organisms emerged. Indeed, the acquisition of multiple signalling pathways was certainly an essential prerequisite for multicellularity. Prokaryotes also express complex cellular communication systems, as will be discussed below.
1.1 Bacterial quorum sensing
Quorum sensing – an example of ancient cellular communication and a forerunner of signalling in multicellular organisms
It is generally believed that cellular communication systems evolved very early in the history of life on Earth. The ability to sense their environment and to communicate with each other would have provided primordial cells with distinct adaptive and replicative advantages. An example of an ancient cellular communication system is quorum sensing, which is observed in both bacteria and archaea and is therefore believed to have originated ~3 billion years ago, well before prokaryotic and eukaryotic life diverged.
Quorum sensing coordinates the behaviour of bacteria within a colony. In particular, quorum sensing is important for controlling bacterial gene expression and maintaining the viability of a bacterial colony. Under certain conditions, bacteria secrete peptides known as autoinducers into their surroundings; if the bacterial population density increases so will the concentration of autoinducer. If the concentration of autoinducer reaches a sufficient level, it can trigger the transcription of specific genes within all the cells in a bacterial colony so that they act in concert (Figure 3). Quorum sensing therefore allows planktonic (single-celled, free-swimming) bacteria to adopt group behaviours when they reach a certain density. The production of autoinducer by bacteria does not have a simple proportional relationship to bacterial population density. Rather, the release of autoinducer substantially increases at a critical population level (dashed vertical line in Figure 3). This surge in autoinducer release underlies the coordinated change from planktonic to group behaviour. Cellular signalling systems often show rapid changes in activity beyond a threshold point. This allows for cellular processes to be either switched on or off with only a relatively small change in stimulation.
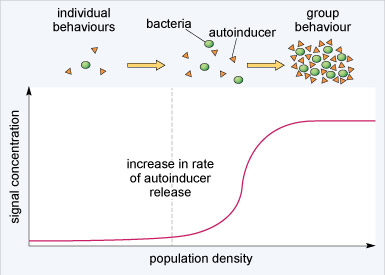
Through changes in the expression of specific genes, autoinducers help bacterial colonies to survive. Typically, autoinducers increase the transcription of genes that encode for antibiotic resistance, in addition to genes that lead to the development of biofilms (Figure 4). A biofilm is a bacterial niche that arises from the aggregation of bacteria within a polysaccharide matrix that they secrete. The bacteria within a biofilm are ~1000-fold less sensitive to antibiotics than their planktonic counterparts and are difficult to kill with disinfectants. Biofilms are found in many places including water distribution pipes and on teeth, lung and intestines. The formation of the protective biofilm niches through quorum sensing is believed to be a major contributor to antibiotic resistance.
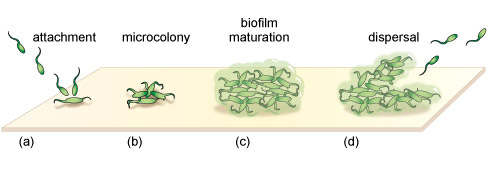
Activity 1 Signalling mechanisms
Figure 5 shows one of the signalling mechanisms that bacteria use for quorum sensing. Describe this quorum sensing mechanism in terms of the generalised cell signalling pathway discussed earlier and represented in Figure 1.
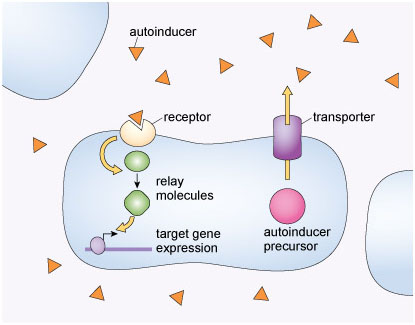
Answer
The bacterial cells release a signalling molecule into their environment (the autoinducer). The signalling molecule binds to specific receptors that are located on the surface of the bacterial cells. The binding triggers a transduction process (involving relay molecules), which activates a response (gene transcription).
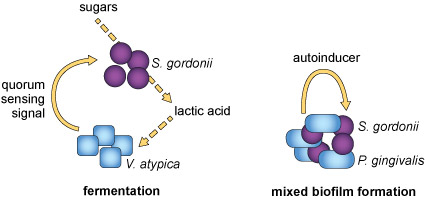
It is believed that the evolution of quorum sensing provided bacteria with a mechanism to coordinate their behaviour. In addition, the effectiveness of autoinducers can tell bacteria something about the environment they live in, such as its viscosity or chemical composition. Moreover, it has been found that different bacterial species can live synergistically within a single biofilm and can signal to each other using quorum sensing molecules, such as autoinducers (Figure 6). These bacterial niches allow interspecies cooperation that is similar to intercellular communication in multicellular organisms.
1.2 Summary of Section 1
- To survive and adapt, cells need to both sense their environment and communicate with each other.
- An important mechanism of cellular communication is the detection of extrinsic signals by specific receptors expressed on cell membranes. This mechanism of cellular communication is used by both prokaryotic and eukaryotic cells.
- Receptors trigger transduction events that relay information within cells. How a cell will respond to an extrinsic stimulus depends on the type of cell, the particular receptors it expresses on its cell membrane and its status.
- Quorum sensing is an ancient form of cellular communication that enables bacteria to coordinate their activities and live in a biofilm niche where they can proliferate.
2 General principles of signalling pathways
Before considering cellular communication systems in detail, it is necessary to discuss some of the general principles by which signalling pathways function.
Cellular communication processes can be described in terms of a series of four fundamental steps.
Activity 2 Cellular communication processes
Drag and drop the descriptions below into the correct sequence to describe a generalised cellular communication process. You can also click or tap through the descriptions in each empty field to make your choices.
Sometimes the extrinsic signal in a cellular communication process is referred to as the ‘first messenger’ and the molecules that convey information within a cell are called ‘second messengers’. While these definitions are commonly encountered in textbooks and the scientific literature, they are often not helpful because signalling typically involves many component factors where designation of a ‘second messenger’ is not appropriate. In addition, the ‘first’ and ‘second’ messengers can sometimes change places. An example of this is calcium, an ion that cells use to control many cellular processes. Calcium ions are released into the cytosol of cells in response to certain extrinsic stimuli. In that context, they can be considered as a second messenger. However, to terminate signalling, the calcium ions within a cell are extruded across the cell membrane. When released into the extracellular space, these calcium ions can bind to calcium-sensing receptors on neighbouring cells and thereby become an extrinsic stimulus. When this happens they could also be considered a first messenger too.
We will avoid use of terms such as first or second messengers, and instead consider cellular communication pathways as sequential chains of interacting components. To understand information flow through signalling pathways, the components are usually considered to be upstream or downstream of other constituents in the pathway. Ultimately, signalling molecules affect the activity of target effector proteins resulting in the cellular response(s). Once a cellular response has taken place, mechanisms that lead to termination of signalling take over. Alternatively, cells can display adaptation or desensitisation; situations where cellular communication is still active, but it causes a lesser effect.
2.1 Summary of Section 2
- Cellular communication is a stepwise process that involves the generation of an extrinsic signal, detection of the signal by a receptor, transduction of the signal by intracellular signalling molecules and a cellular response.
- When the extrinsic signal is removed, cellular communication processes cease.
3 Proteins: major components of signalling pathways
Many signalling pathways, though by no means all, are composed entirely of proteins. To help you make sense of how signalling pathways work, some general principles governing proteins involved in signalling, their functions and their regulation, are reviewed here.
Activity 3 Proteins
Why do you think proteins are particularly well suited for signal transduction?
Answer
Proteins are capable of interacting in a highly specific manner with various ligands and also other proteins. These features are required to ensure fidelity in a signalling pathway. Also, the activity of proteins can be acutely modulated by altering their conformation (for example, by allosteric regulation and by covalent modification), thereby tuning the flow of information via signalling pathways.
Activity 4 Protein–ligand interactions
What is the basis of the specificity exhibited by proteins in protein–ligand interactions?
Answer
If you were thinking along the lines of the structure or conformation of the protein and its ligand, you would be on the right track. Protein–ligand interactions depend on the chemical and physical compatibility of the protein and its ligand and involve the formation of a variety of non-covalent interactions.
Activity 5 non-covalent interactions
Which of the following are non-covalent interactions?
- hydrogen bond
- ionic bond
- hydrophobic interaction
Answer
All of these are types of non-covalent interaction and they are all involved in protein–ligand interactions.
3.1 Signalling proteins act as molecular switches
One of the most important aspects of how proteins function in signalling pathways is that they can act as molecular switches. Most signalling proteins exist in interchangeable active or inactive states. They can therefore behave as rapid molecular switches, being either ‘on’ or ‘off’. Such acute changes in activity are essential for the transmission of signals within cells.
Altered protein expression often accompanies activation of cellular communication for minutes or hours, but changes in protein levels are usually not sufficiently rapid to convey dynamic signals. Rather, the activity of signalling proteins is acutely modulated over a much shorter time frame (seconds to minutes). The most rapid responses are seen with ion channels, where the binding of a ligand causes the channels to open within fractions of a second. The rapid activation of ion channels is critical for the transmission of electrical signals in neurons and muscle cells.
How does a protein actually convey a signal? The answer lies in the conformation of the protein, which is related to its activity. Usually, the upstream signal induces a change in a protein’s conformation, which enables it to carry out its downstream signalling function. Common ways of modulating a protein’s activity are by covalent modification and allosteric regulation.
One of the most commonly encountered forms of covalent protein modification in signalling pathways is phosphorylation. Protein kinases catalyse the transfer of the terminal (γ) phosphate of ATP to a tyrosine, serine or threonine residue on their target protein. Kinases are commonly named in relation to their upstream activator or downstream substrate protein, but are also noted by the amino acids that they phosphorylate. An example of a kinase that you will encounter later in this course is MAP kinase, which is a serine/threonine kinase (often abbreviated to Ser/Thr kinase). You should remember that Ser/Thr kinases phosphorylate serine and/or threonine residues, tyrosine kinases phosphorylate tyrosine residues, whereas dual-specificity kinases can phosphorylate serine, threonine and tyrosine residues. Some kinases actually phosphorylate themselves. This is known as autophosphorylation, and can be critical for kinases to become fully active.
The length of time that a signalling protein remains in its phosphorylated state before being dephosphorylated can be important in determining the signalling outcome. If phosphorylation induces activation, the longer a signalling protein is active, the more downstream signalling molecules it can activate or generate. Many phosphorylated signalling proteins are protein kinases themselves, whose activation results in a series of phosphorylation cascades.
Activity 6 Chemical structures
Proteins can be phosphorylated on serine, threonine or tyrosine residues. The chemical structures of the side chains of these amino acids are shown in Figure 7. What do these all have in common?
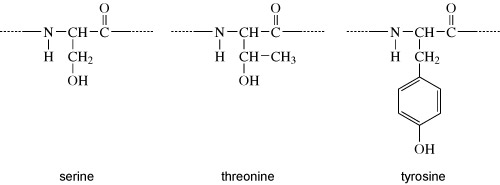
Answer
They all have a hydroxyl group (OH). It is at this hydroxyl group that the phosphate group is added, as shown in Figure 8.
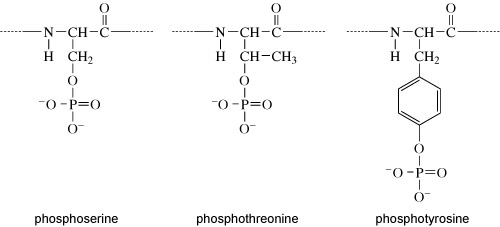
Kinases recognise specific binding sites on their substrate proteins and typically only phosphorylate amino acids within a particular sequence, called a consensus sequence. How does phosphorylation of a protein affect its activity? Addition of a phosphate group can change the conformation of a protein, or alter the interactions of a protein with substrates or other molecules. Some proteins have binding sites for specific phosphorylated motifs on other proteins. Phosphorylation can therefore facilitate the association of proteins. Over a longer timescale, phosphorylation may be a signal for protein degradation, or translocation of a protein to another part of the cell. A phosphate group is removed from a protein by a phosphatase enzyme, generating inorganic phosphate and returning the protein to its unphosphorylated form (Figure 9(a)).
There are numerous different protein kinases in eukaryotic cells. They phosphorylate specific substrates by recognising consensus sequences. Though less numerous than kinases, there are also many phosphatases in eukaryotic cells. Some phosphatases are highly substrate-specific, acting on only one or two phosphorylated proteins, but there are others that can act on a broad range of substrates.
Allosteric regulation of signalling proteins occurs when they bind to molecules including other proteins, lipids or small non-protein ligands including calcium, cyclic AMP and guanosine nucleotides (e.g. GTP). The binding of these allosteric regulators is not random, but is conveyed by specific domains (regions associated with particular functions) within proteins. The binding of an allosteric regulator changes the conformation of its target protein, thereby causing a switch in activity.
Many signalling pathways include guanine nucleotide-binding proteins (G-proteins). These proteins can bind both GDP and GTP, but are only active when they have a GTP molecule bound (Figure 9(b)). There are two classes of G-proteins – trimeric and monomeric. As their names suggest, trimeric G-proteins are composed of three subunits (α, β and γ). Monomeric G-proteins consist of only a single polypeptide that is analogous to the α subunit of trimeric G-proteins.
Trimeric G-proteins interact with a class of receptor known as G-protein-coupled receptors (GPCRs). In their inactive states, both trimeric and monomeric G-proteins have GDP bound. The activation of G-proteins involves the exchange of GDP for GTP. This exchange is promoted by an upstream signalling protein called a guanine nucleotide-exchange factor (GEF). In the case of trimeric G-proteins, the upstream GEF would be a GPCR. Monomeric G-proteins are activated by a range of GEFs that participate in many different signalling pathways. When G-proteins bind GTP they adopt a conformation in which they are active.
G-proteins have the intrinsic ability to hydrolyse GTP to GDP. This means that they are only active for a limited time, because when the bound GTP becomes hydrolysed to GDP they will return to an inactive state. The GTPase activity of G-proteins can be accelerated when they bind to their downstream targets and also by association with regulatory proteins known as GTPase-activating proteins (GAPs).
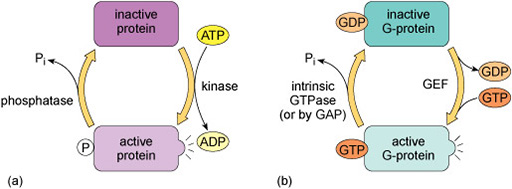
The regulation of G-proteins through binding of guanosine nucleotides is analogous to protein phosphorylation in many ways (Figure 9). In a sense, GEFs play a similar role to protein kinases in that they lead to the activation of signalling; while GAPs are comparable to protein phosphatases that terminate signalling. Moreover, just as the length of time that a protein is phosphorylated determines how long it participates in signalling processes, so the duration of G-protein signalling depends on how long the associated GTP molecule persists before it is hydrolysed to GDP.
3.2 Proteins perform a variety of functions in cell signalling
The proteins involved in signalling can be roughly grouped into a number of categories, based on their functions and activities. Note, however, that the designation of proteins in these categories is not always simple because many proteins within signalling pathways can have more than one function. Some of the ways in which signalling proteins function in signalling pathways are summarised below.
Box 1 Functions of proteins in cell signalling
Relay proteins: These pass a signal on to the next member in the chain. Many different proteins fulfil this role.
Amplifier proteins: These increase the strength of signalling. To achieve this, they either activate multiple downstream targets, or generate large numbers of messenger molecules. Amplifier proteins are often enzymes, such as kinases, or ion channels that can transport many ions in response to a single activation signal.
Transducer proteins: These change a signal into a different form. For example, a kinase might be switched on by allosteric interaction with an upstream molecule, but subsequently activate a downstream protein via phosphorylation.
Bifurcation proteins: These branch a signal to different signalling pathways.
Integrator proteins: These receive signals from different pathways and integrate their input into a common signalling pathway.
Modulator proteins: These regulate the activity of proteins within a signalling pathway.
Anchoring proteins: These tether members of the signalling pathway in particular subcellular locations, such as the cell membrane or the cytoskeleton, thereby ensuring that the signal is being relayed to the right place.
Adaptor proteins: These link one signalling protein with another. Adaptor proteins usually do not act as a signal themselves, but link signalling proteins together so that information can be relayed along a pathway.
Scaffolding proteins: These bind several signalling proteins and may also tether them, forming a much more efficient functional complex. Scaffolding proteins may therefore share attributes of both anchoring and adaptor proteins.
Based on the descriptions of different signalling protein functions in Box 1, attempt to interpret the roles of the proteins in a hypothetical signal transduction cascade in the activity below.
Activity 7 Signal transduction cascades
Consider the hypothetical signal transduction cascade depicted in Figure 10. Different types of signalling protein are listed to the right of the diagram. Drag and drop the labels for protein types into their correct positions on the figure. (You can also scroll through the options and make your choices by repeatedly clicking on or tapping a label field on the figure.)
Note: You can obtain hints on the role of any of the unlabelled proteins by clicking or tapping on them; the hint is displayed at the top of the figure.
3.3 Functional domains in signalling proteins
Signalling proteins typically have conserved amino acid sequences (domains) that enable them to bind to other proteins, lipids or allosteric regulators. Some commonly encountered protein domains and the motifs or targets that they recognise are shown in Figure 11. The domains are often named because of a specific function. For example, the ‘phosphotyrosine-binding domain’ (usually abbreviated to PTB) enables its host protein to bind to another protein that has a phosphorylated tyrosine residue. Other domains are named because they have some sequence homology with the protein in which the domain was first recognised. For example, ‘Src homology 2 domain’ (SH2) is analogous to a phosphotyrosine-binding domain in the protein Src (pronounced ‘sark’). There are tens of different domains that direct proteins to bind a wide range of targets.
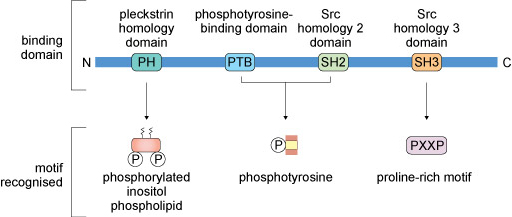
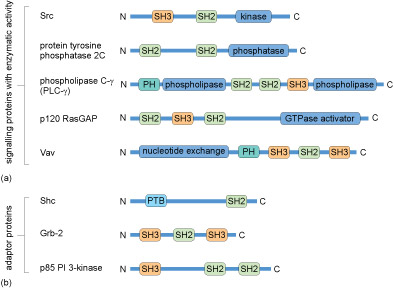
Many proteins contain several of these domains and can bind multiple partners simultaneously. Because of this multiplicity of binding domains, proteins can often form complexes to facilitate the flow of information through a signalling pathway. Linear representations of some signalling proteins and their binding domains are depicted in Figure 12. It is evident that these proteins all have at least two binding domains. All of the different types of proteins listed in Box 1 would have to possess some binding domains; otherwise, they would not be able to interact with upstream or downstream targets.
Activity 8 Protein-binding domains underlie formation of signalling complexes
Figure 13 shows a hypothetical signalling pathway that involves three signalling proteins and an adaptor protein downstream of a receptor; the motifs recognised by particular binding domains are indicated on the figure. Drag and drop the labels onto the unlabelled binding domains to complete the diagram (you can also scroll through the options and make your choices by clicking on or tapping a label field on the figure). You may find it helpful to refer to Figure 11.
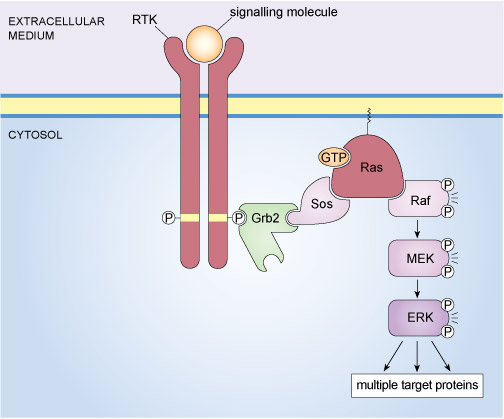
An example of a well-known signalling complex that is formed through specific protein interactions is the mitogen-activated protein kinase (MAP kinase) pathway illustrated in Figure 14. This signalling pathway operates in mammalian cells and is stimulated by the binding of growth factors to tyrosine kinase receptors. The final element of the MAP kinase pathway is a serine/threonine kinase called ERK (an acronym for extracellular signal-regulated kinase); however, there are several steps between the binding of a growth factor to its receptor and the activation of ERK. In essence, a signalling complex comprising a receptor tyrosine kinase (RTK), an adaptor protein (Grb2), a GEF (Sos), a monomeric G-protein (Ras) and three kinases (Raf, MEK and ERK) has to come together at the cell membrane in order for an extrinsic stimulus to cause a cellular response.
There are two isoforms of ERK MAP kinase, referred to as ERK1 and ERK2. Note, however, that since they have similar mechanisms of regulation, scientists don’t always distinguish between these isoforms and use the term ERK to refer to either or both.
The assembly of the components of the mammalian MAP kinase pathway begins with the SH2-containing adaptor protein called Grb2 (growth factor receptor-bound protein 2). Grb2 has no enzymatic activity and consists of little more than two SH3 domains flanking an SH2 domain (Figure 12(b)). Grb2 binds to an activated RTK via its SH2 domain. In turn, Grb2 associates with Sos by virtue of one of its SH3 domains. Sos (which is a GEF) stimulates the exchange of GDP for GTP on the monomeric G-protein Ras, thereby activating it. The active, GTP-bound Ras is then able to recruit the serine/threonine kinase Raf to the membrane, where it becomes activated through a complex series of steps that involves phosphorylation of one Raf molecule by another. Raf subsequently phosphorylates MEK, thus activating it. MEK then phosphorylates the MAP kinase ERK, which goes on to phosphorylate a range of target proteins, including transcription factors.
Signalling through MAP kinases is conserved across the animal and plant kingdoms. A comparison of MAP kinase pathways in various animal species and in plants reveals a similar organisation to the various MAP kinase pathways across species, with the MAP kinase pathways being activated by extrinsic signals that bind to membrane receptors and a cascade of three kinases that relay upstream signals to downstream effectors.
3.4 Summary of Section 3
- Proteins are often the major components of signalling pathways and they have many different roles.
- Proteins can be modified in different ways to control their conformation, activity and interaction with other proteins. These modifications often act as switches to turn signalling processes on or off.
- Kinases phosphorylate other proteins, or themselves, on serine, threonine and tyrosine amino acid residues. Phosphatases remove phosphate groups from proteins. Cells express many different kinds of kinases and phosphatases that have specific protein substrates.
- G-proteins bind GDP or GTP. They are inactive when they have GDP bound, but become active when the GDP is exchanged for GTP. The exchange of GDP for GTP is activated by proteins that function as GEFs (guanine nucleotide-exchange factors). G-proteins hydrolyse GTP to GDP to terminate signalling. The GTPase activity is accelerated by proteins called GAPs (GTPase-activating proteins).
- The ability of a protein to interact with another is determined by conserved protein domains. These are sequences of amino acids that generate an interaction site for one protein to specifically bind to another.
- Many proteins have multiple domains. This allows them to have distinct binding partners and functions.
- Signalling pathways often involve the formation of complexes consisting of multiple proteins, each of which participate in a relay of information from receptor to effector.
Conclusion
In this free course, General principles of cellular communication, you have been introduced to some of the general principles of cellular communication by which extrinsic stimuli are detected by receptors on the surface of cells and, through the activation of specific intracellular signalling pathways, can elicit a response in those cells.
Glossary
- consensus sequence
- A consensus sequence is a common order of nucleotides in DNA, or amino acids in proteins. Proteins interact with DNA, and with other proteins, via specified consensus sequences. In DNA, the consensus sequence is a generalised nucleotide sequence derived from comparisons of closely related DNA sequences from many places within, or between, genomes.
- covalent modification
- The covalent linking to protein amino acid side-chains of groups that can affect their function and/or the localisation of the protein in the cell; such modifications include methylation, acetylation, glycosylation, phosphorylation and lipidation.
- domains
- Physically and functionally distinct regions within a protein molecule.
- G-protein-coupled receptors (GPCRs)
- A family of integral membrane proteins that activate signal transduction processes inside cells; they typically bind extrinsic signals via their carboxy-terminal extracellular domains, and via a conformational change alter the activity of associated GTP-binding proteins. They are sometimes referred to as 7-helix transmembrane (7TM) receptors, serpentine receptors or heptahelical receptors because they possess seven transmembrane domains. GPCRs can also be considered as guanine nucleotide-exchange factors since they promote the exchange of GTP for GDP on associated G-proteins.
- GTPase-activating proteins (GAPs)
- GTPase-activating proteins associate with GTP-binding proteins and accelerate the rate of GTP hydrolysis.
- guanine nucleotide-binding proteins (G-proteins)
- Also known as GTP-binding proteins. Proteins that act as molecular switches, being activated when GTP is exchanged for GDP. Can be monomeric proteins, or exist as a trimeric complex of α, β and γ subunits. With trimeric G-proteins, the α subunit binds guanine nucleotides.
- guanine nucleotide-exchange factor (GEF)
- Proteins that associate with GTP-binding proteins and accelerate the exchange of GTP for GDP to switch on signalling.
- ligand
- A molecule or ion that binds to a receptor or other protein at a specific binding site and may cause activation or inhibition of the receptor and its downstream signalling processes.
- mitogen-activated protein kinase (MAP kinase or MAPK)
- Kinases that were so-named because they were first identified in the signalling pathways activated by molecules that stimulate cells to divide (i.e. mitogens). MAP kinases are components of intracellular signalling cascades that are activated by extrinsic ligands binding to cell membrane receptors; often function in a cascade of kinases in which the final kinase phosphorylates target proteins including transcription factors.
- mitogen-activated protein kinase (MAP kinase) pathway
- Signalling pathways involving MAP kinases.
- molecular switches
- Cellular proteins that change between an active (on) and an inactive (off) conformation. The conformational change is usually brought about by the addition/removal of a phosphate group, either by phosphorylation/dephosphorylation of an amino acid residue or by GTP/GDP exchange.
- motifs
- Sequences of amino acids that are found in a number of related proteins. Alternatively, sequences of bases that are found in functionally related gene segments in different genes, e.g. the target sites for transcription factors. This term is also used to describe combinations of secondary structure (supersecondary structures) in proteins.
- phosphorylation
- The transfer of a phosphate group from one molecule to another. Many proteins are phosphorylated by kinases that transfer the terminal phosphate group from ATP. Phosphorylation changes the conformation of a target protein so that its activity, or ability to interact with other molecules, is altered.
- pleiotropic
- Describes a cellular process, signal or gene that has more than one effect, or more than one phenotypic outcome. At the genetic level, describes a situation where a single mutation affects two or more apparently unrelated phenotypic traits.
References
Further reading
Acknowledgements
This free course was written by Martin Bootman.
Except for third party materials and otherwise stated (see terms and conditions), this content is made available under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 Licence.
The material acknowledged below is Proprietary and used under licence (not subject to Creative Commons Licence). Grateful acknowledgement is made to the following sources for permission to reproduce material in this free course:
Course image: STEVE GSCHMEISSNER/Science Photo Library
Figure 4: Islam, M.S., Richards, J.P. and Ojha, A.K. (2012) ‘Targeting drug tolerance in mycobacteria: a perspective from mycobacterial biofilms’, Expert Review of Anti-infective Therapy Sep 2012, vol. 10, no. 9, pp. 1055–66. Reproduced by permission of Anil Kumar Ojha.
Every effort has been made to contact copyright holders. If any have been inadvertently overlooked the publishers will be pleased to make the necessary arrangements at the first opportunity.
Don't miss out
If reading this text has inspired you to learn more, you may be interested in joining the millions of people who discover our free learning resources and qualifications by visiting The Open University – www.open.edu/ openlearn/ free-courses.
Copyright © 2016 The Open University