The non-coding component of our genome
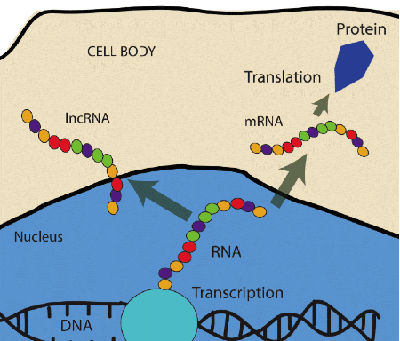 shows the flow of genetic information from DNA to RNA via transcription and then to proteins via translation. As shown some genes do not encode for proteins such as lncRNAs.
Each cell in our body contains all the information to build and maintain our organism alive and active. This information is stored in our genome, which is formed by two long strands of DNA nucleotides creating a double helix structure (fig.1). Our genome contains around 3 billion of nucleotides, which can act like a code to create other molecules called RNAs. This process is called transcription. These RNA transcripts can be used again as a code to create proteins via a process called translation (fig.1). Both transcription and translation work using other molecules called enzymes, factors and cofactors, all of which are proteins with specific functions. Therefore, the classical definition of “gene” refers to DNA sequences, which are transcribed into RNA (mRNA) and then translated into proteins. We know today that some RNAs cannot produce proteins and it has been supposed for many years that they could be useless. We have now evidence that around the 80% of the human genome is transcribed into RNA, but just the 1-2% encodes for proteins. The transcripts lacking the capability to encode for proteins are called non-coding RNAs (ncRNAs). Based on the aforementioned paradigm, it has been supposed for many years that RNAs were “junk genome”. Emerging evidence indicates that ncRNAs play important biological functions, particularly in some diseases. What was considered “junk” emerges today as a gold mine for the discovery of new important findings in biomedical research.
shows the flow of genetic information from DNA to RNA via transcription and then to proteins via translation. As shown some genes do not encode for proteins such as lncRNAs.
Each cell in our body contains all the information to build and maintain our organism alive and active. This information is stored in our genome, which is formed by two long strands of DNA nucleotides creating a double helix structure (fig.1). Our genome contains around 3 billion of nucleotides, which can act like a code to create other molecules called RNAs. This process is called transcription. These RNA transcripts can be used again as a code to create proteins via a process called translation (fig.1). Both transcription and translation work using other molecules called enzymes, factors and cofactors, all of which are proteins with specific functions. Therefore, the classical definition of “gene” refers to DNA sequences, which are transcribed into RNA (mRNA) and then translated into proteins. We know today that some RNAs cannot produce proteins and it has been supposed for many years that they could be useless. We have now evidence that around the 80% of the human genome is transcribed into RNA, but just the 1-2% encodes for proteins. The transcripts lacking the capability to encode for proteins are called non-coding RNAs (ncRNAs). Based on the aforementioned paradigm, it has been supposed for many years that RNAs were “junk genome”. Emerging evidence indicates that ncRNAs play important biological functions, particularly in some diseases. What was considered “junk” emerges today as a gold mine for the discovery of new important findings in biomedical research.
Long non-coding RNAs: how can they act in fundamental biological processes?
It is estimated that the human genome encodes for more than 50,000 non-coding RNAs. Many of these transcripts are longer than 200 nucleotides, and are therefore designated as long non-coding RNAs (lncRNAs) (fig.1). Many lncRNAs are highly expressed in cancer cells, suggesting that they could be involved in fundamental biological processes and could play a role in cancer initiation and progression. Indeed some studies revealed that lncRNAs can act in the nucleus or in the cytoplasm of the cells, thereby regulating cell cycle, cell differentiation and proliferation and acting in the transcriptional regulation of gene expression. The main mechanism of action of lncRNAs seems to be explicated by the interaction with epigenetic effectors. These effectors are protein complexes which can alter our hereditary traits without modifying the DNA sequence. Several studies show that lncRNAs can recruit them to specific region of the DNA where they can turn on and off some genes. While most nuclear lncRNAs control cellular processes through epigenetic regulation, some cytoplasmic lncRNAs can regulate gene functions after their transcription, thereby acting at different levels inside the cell. Since they could act via several mechanisms, it is estimated that increasing or decreasing the production of lncRNAs, through genetic engineering techniques, can influence many physiological and pathological processes, thereby paving the way for the treatment of several diseases.
LncRNAs and Cancer
Studying lncRNAs could have a fundamental importance to shed light on some mechanisms of cancerogenesis and to find new targets for cancer diagnosis, prognosis and therapy. Some lncRNAs are being tested in clinical trials, since they represent some of the most differentially expressed transcripts between primary cancers (where the tumour first started to grow) and metastatic cancers. Metastasis is the mechanisms by which tumour cells can grow in different sites and organs, spreading from the primary site. Even if many lncRNAs have an oncogenic role and promote the acquisition of aggressiveness in some cancers, tumour-suppressive lncRNAs have been described as well, downregulated in aggressive cancers with high metastatic potential. So can lncRNAs be a source of new discoveries in medicine? Can they help us to understand how our organism works? Are some of them really junk material for scientific research? We don`t know it completely yet but a new era of scientific challenges has started.

Rate and Review
Rate this article
Review this article
Log into OpenLearn to leave reviews and join in the conversation.
Article reviews