1.6 Signalling proteins can act as molecular switches
How does a signalling molecule actually convey a signal? With second messengers, it is easy to understand: they are produced or released in large quantities, diffuse to their target, to which they usually bind, bringing about a functional change, after which they are degraded or stored within a subcellular compartment (such as endoplasmic reticulum). With signalling proteins it is less obvious. Protein concentrations cannot fluctuate rapidly, and protein molecules cannot easily move within the cell. The conformation of many proteins is related to their activity, and is subject to regulatory mechanisms.
What are the mechanisms by which proteins can be switched from one conformation to another?
One way of modulating a protein's activity is by allosteric regulation, whereby binding of a small ligand induces a conformational change in the protein. Another way is by addition of a negatively charged phosphate group, either by phosphorylation of an amino acid residue by a protein kinase or by binding of a GTP molecule instead of a GDP (G proteins).
Although allosteric regulation by binding small molecules is a widespread regulatory mechanism for the activity of many proteins, including receptors and structural, motor and signalling proteins, the addition or loss of phosphate groups usually drives most functional changes in the sequence of activation/deactivation steps that form a typical intracellular signalling pathway. In reality, many intracellular signalling proteins act as molecular switches. What often happens is that the proteins can be temporarily modified, converting them from an inactive (non-signalling) form to an active (signalling) form (Figure 2), or vice versa. Usually the upstream signal induces a change in the protein's conformation, which enables it to carry out its downstream signalling function. The reason why such molecules are sometimes referred to as molecular switches is because they are either ‘on’ or ‘off’. These proteins can be grouped according to how they are switched on/off, rather than their subsequent mode of action. As outlined above and in Figure 8, signalling molecular switches mainly belong to two categories.
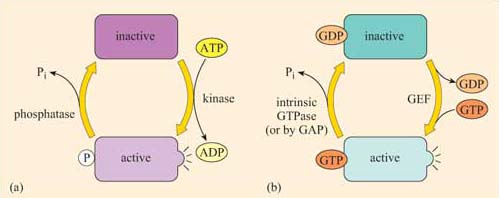
One group of proteins often encountered in signalling are those that are modified by phosphorylation of an amino acid residue by an upstream kinase (Figure 8a). The phosphate is derived from the terminal (γ) phosphate of ATP, and added covalently to a tyrosine, serine or threonine residue by a protein kinase. Phosphorylation usually, but not necessarily, activates a protein. Sometimes, however, it may cause a conformational change that inactivates the protein.
The phosphate group is subsequently removed by a phosphatase, generating Pi (inorganic phosphate), and the protein reverts to its original form. The length of time that the protein remains in its phosphorylated state before being dephosphorylated can be important in determining the signalling outcome. If phosphorylation induces activation, the longer a signalling protein is active, the more downstream signalling molecules it can activate (or the longer that second messengers are synthesized or released by an active signalling protein, the higher the concentrations that they achieve). It is important to note here that many phosphorylated signalling proteins are protein kinases themselves, whose activation results in a series of phosphorylation cascades, as you will see in Section 3.6 (see also Box 1).
The second main group of signalling molecular switch proteins are the GTP-binding proteins, known as G proteins (Figure 8b). In this case, the on/off state characterized by the addition/loss of a phosphate group is not mediated by covalent binding of a phosphate group, but by the binding of a GTP molecule and its hydrolysis to GDP.
In the same way that the rate of dephosphorylation of a phosphorylated protein determines how long it remains active, the length of time that a GTP-binding protein remains active (and hence the number of downstream molecules it can activate) is determined by the rate of GTPase activity. In a sense, GEFs play a similar role to protein kinases and GAPs are comparable to protein phosphatases. In their active form, G proteins also cause a cascade of phosphorylation events, ultimately resulting in a cellular response.
Box 1 Identification of phosphorylated residues in signalling proteins
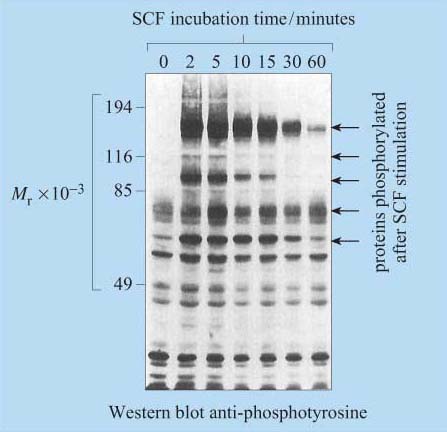
For many years, phosphopeptide and phosphoamino acid mapping has been a useful method used for identifying protein phosphorylation sites. Cells are metabolically labelled with radioactive Pi, and protein extracts are subjected to polyacrylamide gel electrophoresis (SDS–PAGE and Western-blotted onto a special membrane. The protein of interest is then isolated, hydrolysed into peptide fragments by proteases or into individual amino acids by hydrochloric acid, and are then separated by two-dimensional thin-layer chromatography on cellulose plates. The extent of phosphorylation of tyrosine, threonine and serine residues is finally established by autoradiography. Another technique, first developed in the 1980s, involves the use of monoclonal anti-phosphotyrosine antibodies, which specifically recognize phosphorylated tyrosine residues in many proteins. For investigation of signal transduction mechanisms, this was an essential tool for studying the activity of tyrosine kinases and phosphatases. The antibody can either be used to probe Western-blotted proteins (Figure 9) or, in a more refined technique, can be used to immunoprecipitate the phosphoproteins before separating them by SDS–PAGE.
However, these techniques require the use of populations of single cell types, as these antibodies would not differentiate between cell types in mixed cell populations.
More recently, other polyclonal and monoclonal antibodies targeted to phosphorylated residues (serine, threonine and/or tyrosine) within a specific amino acid sequence of a protein have been developed. For example, there are antibodies that recognize phosphorylated Tyr 527 of Src, and others that recognize Tyr 416 of Src, providing a rapid and easy experimental methodology for the study of Src activation. The use of antibodies specific for phosphorylated amino acid residues has allowed the study of signalling protein activation in vivo on tissue sections using immunocytochemical techniques. Cocktails of 30 or more of these antibodies can also be used in combination to simultaneously detect the activation state of several signalling pathways by probing proteins separated in 2-D gels.
Molecular switches can be a lot more sophisticated than a single on/off function. A protein can be phosphorylated at multiple sites, which may have different effects on its activity.
Integrate many different signals such that the signalling outcome is determined by the summation of signalling inputs. Therefore, they behave as specific signal integrators.