Use 'Print preview' to check the number of pages and printer settings.
Print functionality varies between browsers.
Printable page generated Tuesday, 21 May 2024, 5:31 PM
Introducing antimicrobial resistance
Introduction
This module will introduce how antibiotics work and how bacteria acquire and transmit resistance.
You will start by exploring how antibiotics can exert powerful antibacterial effects but be generally well tolerated by people and animals, focusing on the different modes of antibiotic action. You will then consider how bacteria develop resistance in order to protect themselves from antibiotics. You will explore the main mechanisms of antibiotic resistance as well as the differences between intrinsic and acquired resistance, including how acquired resistance is transferred.
You may be familiar with the terms ‘antimicrobial’, ‘antibacterial’ and ‘antibiotic’. Each has a slightly different meaning, but in this module, we will use the term ‘antibiotic’ to refer to any drug that is active against bacteria.
After completing this module, you will be able to:
- explain how antibiotics work against bacterial pathogens
- state what is meant by the term ‘antibiotic resistance’
- explain that antibiotic resistance is a natural phenomenon that protects bacteria from hostile environments
- explain how resistance mechanisms arise, and how resistance can be passed on during bacterial replication
- describe the horizontal gene transfer mechanisms that allow antibiotic resistance to be transferred between bacteria
- apply scientific terminology when explaining how antibiotic resistance relates to your current work.
Activity 1: Assessing your knowledge and skills
Before you begin this module you should take a moment to think about the learning outcomes and how confident you feel about your knowledge and skills in these areas. Do not worry if you do not feel very confident in some skills – they may be areas that you are hoping to develop by studying these modules.
Now use the interactive tool to rate your confidence in these areas using the following scale:
- 5 Very confident
- 4 Confident
- 3 Neither confident nor not confident
- 2 Not very confident
- 1 Not at all confident
Try to use the full range of ratings shown above to rate yourself:
Although this is an introductory course on antibiotic resistance, it assumes that you have a basic understanding of DNA and proteins. If you are unfamiliar with these concepts, you might want to try our free OpenLearn course What do genes do? and listen to our set of audios on DNA, RNA and protein formation before you begin this course.
Activity 2: New terms from Introducing antimicrobial resistance
As you work through the module, put together a personal list of terms that are new to you. Make sure that you feel confident with what they mean and that you can apply them to your own working practice.
- Have you heard any of these terms mentioned in your day-to-day work?
- What was the context that they were used in?
- Can you think of how you would use some of these terms in your day-to-day work?
Add your notes to your reflective blog.
1 How do antibiotics work?
1.1 Selective toxicity
Drugs should demonstrate ‘
Underpinning selective toxicity are the differences between bacterial pathogens and human or animal cells. It is these differences that antibiotics (and other drugs) exploit to exert their specific effects.
There are two distinct types of cell. Bacteria are
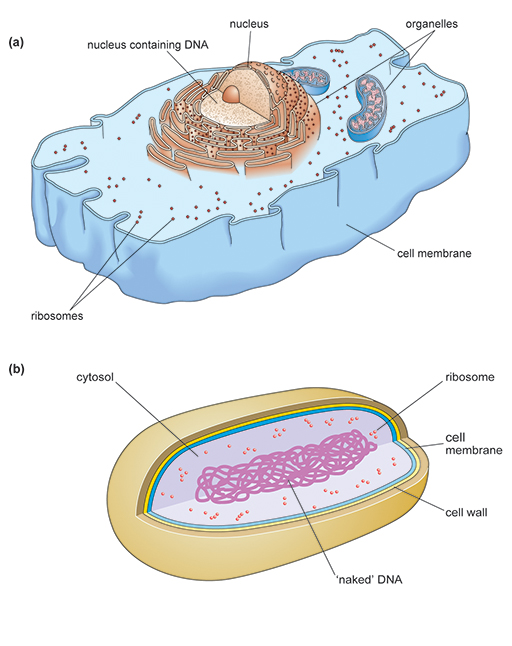
Although there are similarities between the two cell types, eukaryotic cells are structurally much more complex. Prokaryotes and eukaryotes carry out the same essential processes necessary for survival, such as making new proteins,
Antibiotics are selectively toxic because they target structural features or cellular processes in the bacterial pathogen that are different or lacking in the host’s cells.
1.2 Potential bacterial targets for antibiotics
In Activity 3 you will discover which essential cell processes in the bacterial pathogen are potential targets for antibiotics.
Activity 3: When are bacteria vulnerable to antibiotics?
Watch the video about key bacterial cell processes and answer the related questions. You can pause the video to work through this activity at your own pace.
If you are unable to watch this video, or any other video in this course, you can view a transcript of the content by clicking on ‘Show transcript’ below.

Transcript: Video 1 How do antibiotics work?
(a) Suggest a likely consequence for the cell if DNA replication is blocked.
Answer
Blocking DNA replication would impair cell division and kill the bacterial cell.
(b) Which stage or stages of protein synthesis could be targeted by antibiotics?
Answer
Interference with either stage of protein synthesis could result in faulty enzymes and/or structural proteins.
(c) DNA replication, metabolic reactions and protein synthesis also occur in eukaryotic cells. Suggest why antibiotics that target these bacterial processes demonstrate selective toxicity.
Answer
Although cellular processes of prokaryotic and eukaryotic cells have many similarities, antibiotics are selected for clinical use that target those processes that are wholly or partly unique to the bacterial pathogen. This minimises the risk of side effects in the patient.
(d) What might happen to a cell that can no longer make a cell wall?
Answer
Bacterial cells that lack a cell wall are in danger of bursting if too much water moves into the cell by osmosis
(e) Why do antibiotics that target cell wall synthesis leave human and animal cells unharmed?
Answer
Human and animal cells lack a cell wall.
(f) A relatively small number of antibiotics target the bacterial cell membrane. Such antibiotics are often highly toxic to the host. Can you suggest a reason for this?
Answer
The membrane of animal and human cells has a very similar structure to that of bacteria. The potential for such antibiotics to adversely affect eukaryotic cells is therefore greater and these antibiotics generally demonstrate poor selective toxicity. This increases the risk of harmful side effects for the patient.
1.3 Types of antibiotic
Antibiotics may be active against a wide range of bacteria (
Factors that determine the spectrum of antibiotic activity include:
- ability to penetrate the bacterial cell – since most bacterial targets are located in the cell’s interior
- how widespread the target is among different bacterial species
- bacterial resistance to the antibiotic.
1.3.1 Gram-positive and Gram-negative bacteria
Can you remember the importance of the cell wall to bacterial cells?
It is a protective outer layer that provides mechanical support to the cell. It also prevents harmful surges of water moving into the cell by osmosis, which could cause it to burst (lysis).
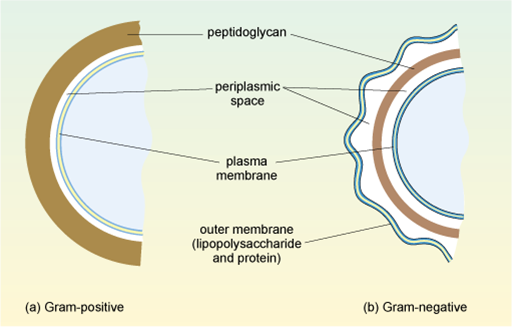
Bacteria are divided into two main groups based on how the cell wall appears when they are stained using Gram straining. This procedure allows the composition of the wall to be visualised. Some ‘atypical’ bacteria, such as Mycoplasma and Chlamydia, cannot be seen by Gram staining and so cannot be categorised in this way, but these will not be considered in this module.
In
In
(For more information on the Gram stain and other techniques used in the laboratory to help identify bacteria you could visit the module entitled Isolating and identifying bacteria.)
This second, outer membrane of Gram-negative bacteria is an effective barrier, regulating the passage of large molecules such as antibiotics into the cell. In contrast, the thick, porous peptidoglycan layer in the cell wall of Gram-positive bacteria gives greater access to antibiotics, allowing them to more easily penetrate the cell and/or interact with the peptidoglycan itself.
1.3.2 Activity against Gram-positive and Gram-negative bacteria
Narrow-spectrum antibiotics are effective against either Gram-positive or Gram-negative bacteria, whereas broad-spectrum antibiotics are effective against both types.
Not all Gram-positive and/or Gram-negative bacteria are affected by a single antibiotic. Can you suggest a possible reason for this?
This is because of the specificity of the antibiotic/bacterial target interaction, whether the bacterial species has the target in question and whether the bacteria are resistant to the antibiotic.
1.3.3 Bactericidal versus bacteriostatic antibiotics
While some antibiotic classes have consistent antibacterial effects, such as β-lactams which are nearly always bactericidal, the activity of other classes may depend on the dose of antibiotic prescribed or how long the treatment lasts. For example, fluoroquinolones and aminoglycosides, while usually bactericidal, may be bacteriostatic when used at low concentration.
You should by now have a good idea of how antibiotics interact with bacterial cells. You will now consider what happens to the bacterial population as a whole when antibiotics are administered.
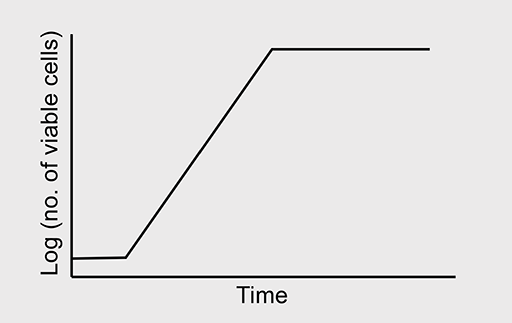
In nature, bacterial growth follows a typical pattern shown in Figure 3. The growth curve comprises four phases:
- The lag phase: Bacteria are adapting to their environment; nutrients are plentiful and the cells grow in size. Cell number remains relatively constant, balanced by the deaths of some cells and division of others.
- The exponential/logarithmic (log) phase: This phase marks a big increase in cell numbers. Maximum growth rate is achieved, with a constant doubling of the bacterial population. Growth then slows as nutrients become depleted and bacterial waste products build up to toxic levels.
- The stationary phase: The bacteria enter this phase when the number of new cells equals the number of cells dying. The total number of cells in the population remains constant.
- The death/decline phase: Unless nutrients are replenished and waste products are removed, the bacteria progress to the death phase. More cells die than are produced and the number of cells in the population declines; however, some cells may remain viable (capable of surviving).
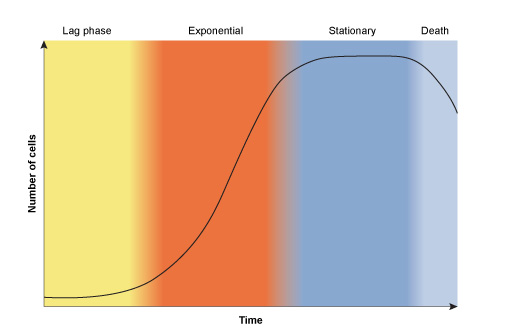
Bacteria are at their most susceptible to antibiotic attack when they are actively growing. You will now consider what happens to a bacterial culture when antibiotics are introduced during this exponential phase of growth.
Activity 4: Effect of antibiotics on bacterial growth
Bacteriostatic antibiotics
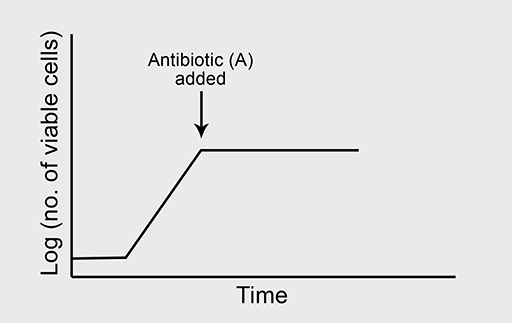
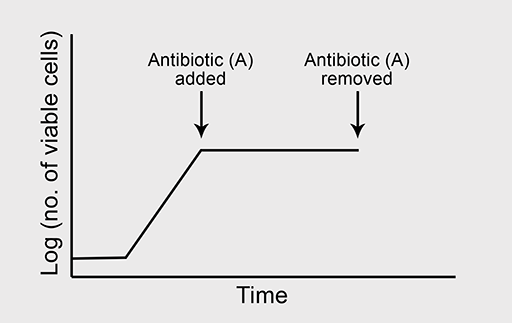
(a) A typical growth curve is shown in Figure 4a.
Figure 4b shows the normal growth curve of a bacterium which is sensitive to the bacteriostatic antibiotic ‘A’. What will happen to the bacterial growth rate when ‘A’ is added to the culture in high concentration where all other growth conditions are optimal?
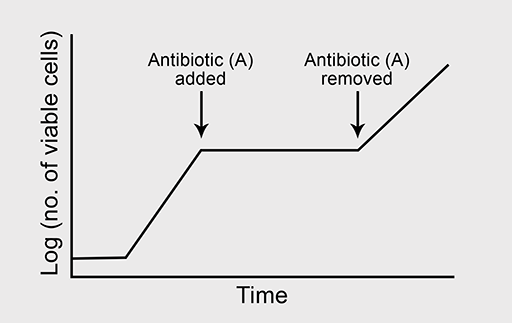
(b) Now predict what will happen to bacterial growth if antibiotic A is removed from the culture at the point indicated on the graph in Figure 4d.
Bactericidal antibiotics
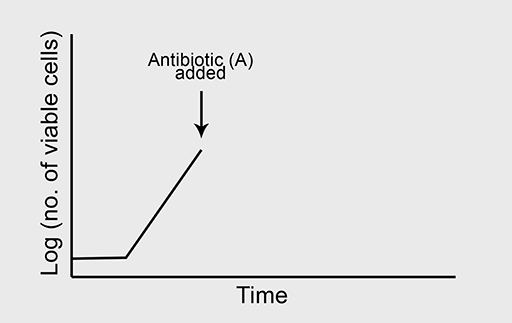
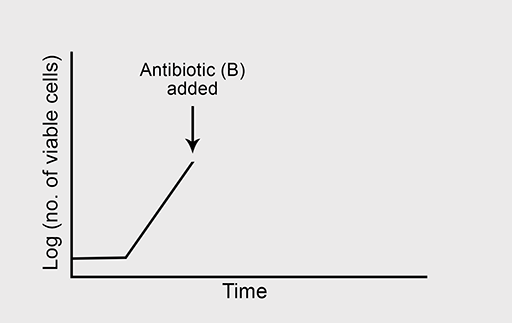
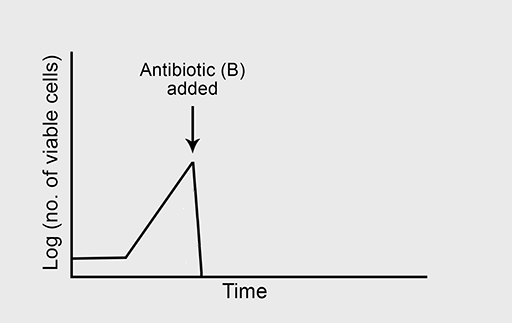
Figure 5a shows the normal growth curve of a bacterium that is sensitive to the bactericidal antibiotic ‘B’. What will happen to the rate of bacterial growth when B is added to the culture in high concentration? Again, you should assume that growth conditions are otherwise optimal.
Answer
Bactericidal antibiotics kill susceptible bacteria during the exponential phase of growth and help to eliminate the infection.
Bacteriostatic antibiotics stop bacterial growth even though the cells remain viable. This allows time for the host’s immune system to be activated and target the bacterial pathogen – again effecting a cure.
1.4 Antibiotic modes of action
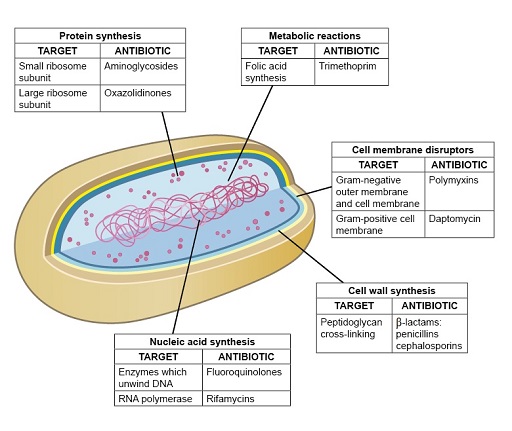
This section focuses on the four main modes of antibiotic action (Figure 6) that lead to inhibition of one of the following:
- cell wall synthesis
- protein synthesis
- nucleic acid synthesis
- metabolic reactions
- cell membrane function.
Don’t worry if you don’t understand all of these terms, as they will be explained in later sections.
Members of the same class of antibiotics share a characteristic structural feature that determines the drug’s affinity and specificity for target molecules in susceptible bacteria. You will now look in more detail at antibiotics that exemplify each of these modes of action.
1.4.1 Inhibitors of cell wall synthesis
As you saw in Activity 3, the cell wall is essential for normal functioning of the bacterial cell. Antibiotic inhibitors of cell wall synthesis block the production of
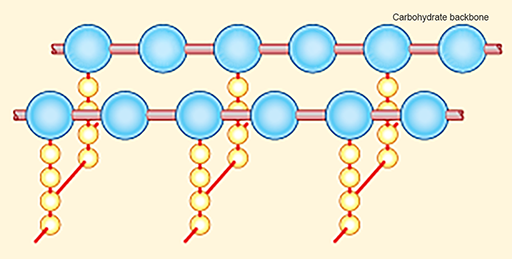
Disruption of the peptidoglycan layer of the cell wall can therefore result in cell lysis (Figure 8).
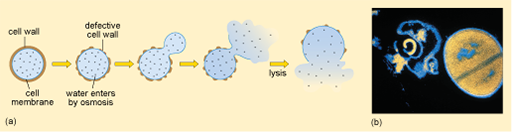
Examples of cell wall synthesis inhibitors are the β-lactam antibiotics. These include penicillin and its derivatives, and the cephalosporins. All β-lactam antibiotics contain a core chemical structure called a β-lactam ring (Figure 9) that determines the mode of action of this class of antibiotics.
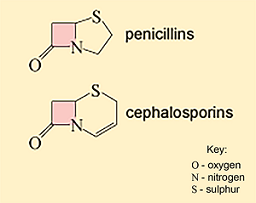
The β-lactam antibiotics interfere with the formation of the peptidoglycan cross-links by inhibiting the enzymes responsible for cross-linking adjacent molecules in the peptidoglycan layer. The β-lactam antibiotics bind to these enzymes, collectively known as penicillin-binding proteins (PBPs) and prevent them from forming cross-links.
Figure 10 shows what happens when a β-lactam antibiotic, in this case penicillin, binds to an active PBP.
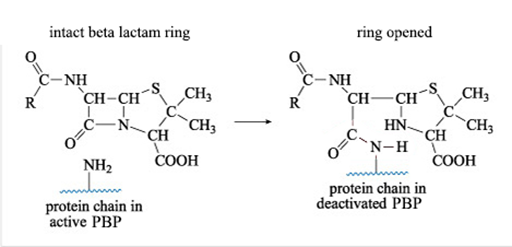
The reaction shown in Figure 10 results in a new PBP side chain which is much larger than the original –NH2 group and effectively deactivates the PBP.
By interfering with the formation of peptidoglycan cross-links β-lactam antibiotics weaken the cell wall. In addition, repairs cannot be made to the cell walls, allowing the osmotic pressure inside the cells to increase, eventually leading to lysis.
Later in the module you will learn more about how disrupting the interaction between β-lactam antibiotics and PBP contributes to antibiotic resistance mechanisms.
The glycopeptides (e.g. vancomycin) are another group of antibiotics that inhibit cell wall synthesis, but by a different mechanism.
1.4.2 Inhibitors of protein synthesis
Cells synthesise new proteins in ribosomes that are made up of one large and one small subunit. These subunits differ structurally and chemically between prokaryotic and eukaryotic ribosomes (Figure 11). These differences mean that antibiotics can target bacterial ribosomes but have minimal effect on the ribosomes of the host cell.
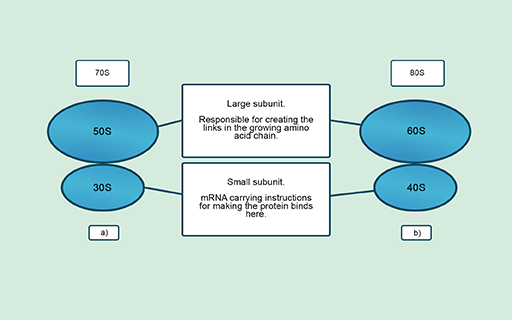
Several antibiotics that you may be familiar with, such as gentamicin, tetracycline and erythromycin, work in this way. Interfering with ribosome functions means that cells cannot produce essential proteins and will either die or be unable to replicate. Other antibiotics, such as fusidic acid, also inhibit the production of proteins by blocking different parts of the protein synthesis pathway.
1.4.3 Inhibitors of nucleic acid synthesis
Differences between enzymes that carry out the synthesis of nucleic acids in eukaryotic and prokaryotic cells allow antibiotics to target these processes in bacteria.
For example, fluoroquinolones such as ciprofloxacin and levofloxacin (Figure 12) specifically inhibit bacterial enzymes that unwind the DNA double helix, separating the two strands so that the DNA can be copied. If the strands of DNA do not unwind and separate, bacterial
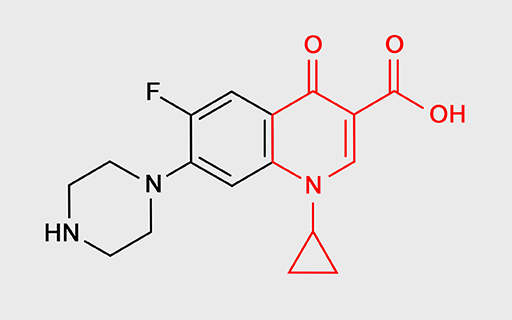
Another class of antibiotics – rifamycins – inhibit RNA synthesis by binding to and inhibiting an enzyme called RNA polymerase. This enzyme transfers the instructions carried by genes to the intermediary molecule, mRNA. Interference in this process ultimately stops new proteins being made. Rifampicin is a commonly used rifamycin.
1.4.4 Inhibitors of metabolic reactions
Antibiotics that disrupt essential bacterial metabolic pathways are acting as
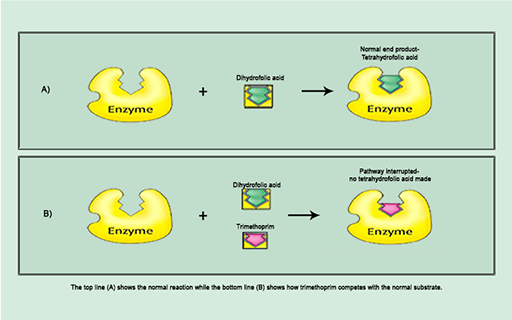
For example, trimethoprim inhibits the synthesis of folic acid, a vitamin that bacteria, unlike humans, must make themselves. Trimethoprim is a

The action of trimethoprim illustrated in Figure 13b exemplifies the specific interaction between antibiotic and bacterial target at a molecular level that disrupts a particular cellular process.
Identify the structure labelled ‘enzyme’.
The enzyme is dihydrofolate reductase.
1.4.5 Inhibitors of cell membrane function
The cell membrane controls the passage of substances into and out of the cell. Inhibitors of the cell membrane function interfere with the cell membrane’s structure, causing cell contents to leak out, and eventually leading to cell death.
Daptomycin is a cell membrane inhibitor that is used to target Gram-positive bacteria. You may recall that Gram-positive cells have a large peptidoglycan cell wall, covering the plasma membrane. Daptomycin binds with calcium ions to form a calcium complex. The addition of the calcium to form the calcium complex means that the daptomycin can insert itself into the plasma membrane. The complexes then aggregate within the plasma membrane to form pore-like structures that allow ions to leak out of the cell, eventually leading to cell death (Figure 14). Daptomycin is a relatively new antibiotic and is not in widespread use, which may explain why resistance, for now, is rare.
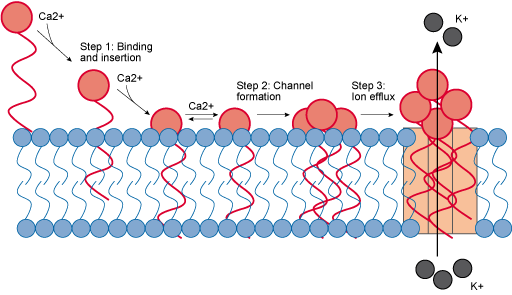
Polymyxins, such as colistin, are another group of cell membrane inhibitors that are used to target Gram-negative cells. Remember, the Gram-negative cell has an additional outer membrane that is not present in Gram-positive cells. Polymyxins bind to the lipopolysaccharides within the outer membrane, causing structural changes to the membrane. This in turn causes a loss of membrane integrity, increasing its permeability. The polymyxins are now able to pass through the proteoglycan wall and reach the plasma membrane, disrupting it in a similar way. This causes the cell contents to leak out, eventually leading to cell death.
2 How do bacteria become resistant to antibiotics?
The ability to divide rapidly and reproduce exponentially means that bacteria are constantly mutating and adapting to survive any changes in their environment. Antibiotic resistance is the ability of bacteria to resist the action of antibiotics so that they survive exposure to antibiotics that would normally kill them or stop their growth (CDC, 2017; PHE, 2017).
You could be forgiven for thinking that antibiotic resistance is only caused by human use, and misuse, of antibiotics. However, as you will see in the following video, bacteria that have not been exposed to antibiotics used by humans have acquired resistance to many of the antibiotics we use to treat infections; as you will see later, some may be inherently resistant if they lack the pathway or component that the antibiotic acts on: for example, colistin has no effect on Gram-positive organisms, because they do not have an outer membrane. This video also appears in The problem of AMR, where it was included to show the ubiquity of antimicrobial resistant bacteria. We include it again here so that you can begin to think about the mechanisms that give rise to AMR, which you’ll learn about below.

Transcript: Video 2 Antibiotic resistance is a natural bacterial defence mechanism.
As you may now realise, many antibiotics in clinical use have originated from substances used by micro-organisms to defend themselves against other micro-organisms. In essence, humans have taken these important molecules and made use of them to produce the antibiotics we use. It therefore makes sense that bacteria would derive ways to defend themselves from antibiotics.
2.1 Antibiotic resistance mechanisms
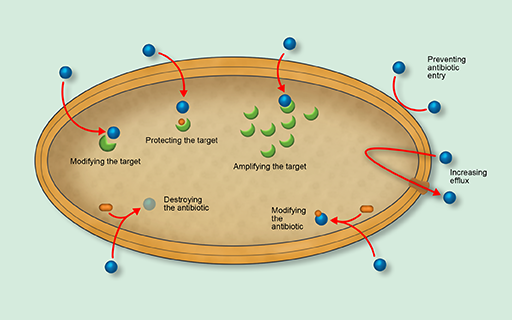
Bacteria have evolved several sophisticated antibiotic resistance mechanisms. Figure 15 gives an overview of the major mechanisms by which bacteria become resistant to the action of antibiotics. Don’t worry if you don’t understand all of these terms, as they will be explained in the following sections.
In this section you will look at the three main mechanisms of antibiotic resistance:
- modifying the antibiotic target
- destroying or modifying the antibiotic
- preventing the antibiotic from reaching its target.
Although you will look at each of these mechanisms in turn, it is worth remembering that bacteria may use multiple resistance strategies simultaneously to survive exposure to antibiotics.
2.1.1 Modifying the antibiotic target
As you already know, antibiotics are selectively toxic because they target structural features or cellular processes in the bacterial pathogen that are different or lacking in the host’s cells. Recall how penicillin and other related β-lactam antibiotics work by binding to penicillin-binding proteins (PBPs), preventing them from binding to their normal target, peptidoglycan. Or how trimethoprim prevents dihydrofolate reductase reacting with dihydrofolic acid.
Changes to the structure of the target that prevent efficient antibiotic binding but still enable the target to carry out its normal function will confer antibiotic resistance (Figure 16).
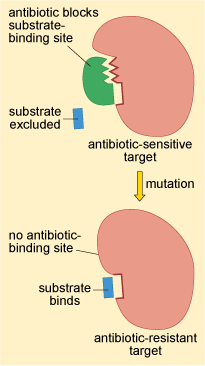
This resistance strategy is widespread among bacteria. Altering the structure of the PBP, for example, is one way that bacteria develop resistance to penicillin.
2.1.2 Destroying or modifying the antibiotic molecule
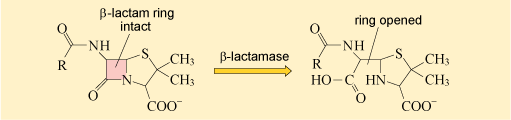
The second mechanism of antibiotic resistance you will look at is the destruction or modification of the antibiotic by bacterial enzymes. Probably the most well studied example of enzymes that destroy antibiotics are the β-lactamases.
As you may recall, β-lactamases deactivate the β-lactam ring of β-lactam antibiotics, preventing them from binding to their target (Figure 17).
The first β-lactamase to be identified was penicillinase. As its name suggests, penicillinase can hydrolyse penicillin but not other, more recently discovered β-lactam antibiotics like cephalosporins. In the 1980s, a new group of β-lactamase enzymes were detected in Europe that hydrolyse a wider range of β-lactams. These enzymes are known as extended spectrum β-lactamase (ESBL).
ESBLs can deactivate almost all of the β-lactam antibiotics currently in therapeutic use. Consequently, their presence significantly reduces the available treatment options for infections caused by bacteria expressing β-lactamase. Since they were first described in the early 1980s, the frequency of infections caused by ESBL-producing bacteria has been increasing. Consequently, ESBLs represent an ever-growing healthcare challenge.
Some β-lactamases can be counteracted by using a β-lactamase inhibitor (a drug that can block the action of the enzyme, such as clavulanic acid or tazobactam) in combination with a β-lactam antibiotic (e.g. clavulanic acid and amoxicillin are combined in co-amoxiclav); however, even these are not necessarily effective against ESBLs, and each combination needs to be tested against the bacteria before using clinically.
How might a β-lactamase inhibitor help the treatment of infections caused by β-lactamase-expressing bacteria?
The β-lactamase inhibitor will block the ability of the β-lactamase to deactivate the β-lactam antibiotic so that the antibiotic can bind to its target molecule.
Other antibiotic-modifying enzymes do not destroy or target the core chemical structure that confers antibacterial activity. Instead they modify the antibiotic’s structure by adding chemical groups that prevent it from binding to its target. One group of antibiotics that are particularly susceptible to modification are the aminoglycoside antibiotics which include streptomycin and gentamicin (Figure 18).
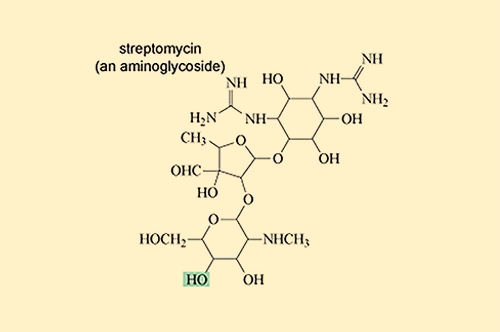
Aminoglycoside-modifying enzymes add bulky chemical groups to the exposed hydroxyl (–OH) and amino (–NH2) groups of the antibiotic, thus preventing it from binding to its target.
2.1.3 Preventing entry, increasing exit
Antibiotics are only effective if they can reach their target. Preventing antibiotics from reaching their target is the final mechanism of antibiotic resistance that you will look at in this module.
As you should recall, the cell wall protects bacteria from osmotic and mechanical damage. To reach their targets inside the cell, antibiotics must cross this cell wall. Bacteria can either modify their cell wall to prevent antibiotics entering the cell or can use special structures, called efflux pumps, to remove antibiotics that have gained entry. In Activity 5 you will look at these mechanisms in more detail.
Activity 5: Transporting antibiotics across the bacterial cell wall
First, watch the following animation which describes how antibiotics are transported across the bacterial cell wall.
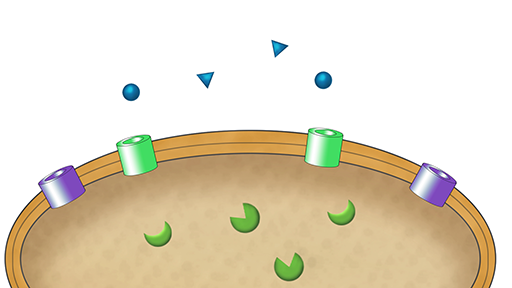
Transcript: Video 3 Animation of the mechanisms of transport of antibiotics across the membrane.
Now answer the following questions.
a.
(a) decreases the amount of antibiotic entering Gram-negative bacteria
b.
(b) increases the amount of antibiotic entering Gram-negative bacteria
c.
(c) has no effect on the amount of antibiotic entering Gram-negative bacteria.
The correct answer is a.
a.
Your answer is correct. Most antibiotics cannot cross the outer membrane of Gram-negative bacteria and therefore enter the cell via porin channels. Decreasing the number of porin channels will decrease the amount of antibiotic entering the bacteria.
a.
(a) very few porin channels on their outer membrane or have replaced their porin channels with channels that exclude fluroquinolone antibiotics
b.
(b) numerous porin channels on their outer membrane
c.
(c) replaced their porin channels with channels that selectively transport fluroquinolone antibiotics.
The correct answer is a.
a.
Your answer is correct. If an antibiotic cannot reach its target, bacteria will be resistant to its action. Decreasing the expression of porins, or replacing them with channels that cannot transport the antibiotic, will prevent the antibiotic from crossing the outer membrane and reaching its target; therefore, these bacteria will be resistant.
a.
(a) increase the amount of fluroquinolone antibiotics in the bacterial cell
b.
(b) decrease the amount of fluroquinolone antibiotics in the bacterial cell
c.
(c) have no effect on the amount of fluroquinolone antibiotics in the bacterial cell.
The correct answer is b.
b.
Your answer is correct. Efflux pumps actively transport antibiotics out of the bacterial cell. Therefore, increasing transport through these channels will decrease the amount of antibiotic inside the cell.
a.
(a) efflux pumps that are unable to transport fluroquinolone antibiotics
b.
(b) efflux pumps that transport fluroquinolone antibiotics
c.
(c) no efflux pumps.
The correct answer is b.
b.
Your answer is correct. If an antibiotic cannot reach its target, bacteria will be resistant to its action. Actively transporting antibiotics out of the cell decreases their concentration inside the cell, so that they cannot build up to a high enough concentration to exert the effect on their target.
Increasing active transport by expressing more efflux pumps that can actively transport the antibiotic out of the cell decreases the amount of antibiotic inside the cell and prevents it from acting on its target.
As you should now appreciate, bacteria can prevent antibiotics from reaching their target by decreasing the permeability of their outer membrane or by actively transporting antibiotics out of the cell (Activity 3). Both decreased porin expression and increased efflux pump expression have been reported in antibiotic-resistant clinical
2.2 Intrinsic and acquired resistance
There are two ways in which bacteria can have these resistance mechanisms:
- intrinsic (or inherent) resistance
- acquired resistance.
In this section, you will look at each type in turn.
2.2.1 Intrinsic resistance
Intrinsic resistance may occur because bacteria lack the target for a particular antibiotic or because the drug can’t get to its target. It reduces the pool of antibiotics available to treat infections.
It is worth noting that resistance elements that are intrinsic to one bacterial type can occasionally be transferred to another one, and you will learn about this below.
2.2.2 Introducing acquired resistance
As its name suggests,
Unlike intrinsic resistance, acquired resistance is only found in some populations of a bacterial type.
Acquired resistance is a very significant healthcare concern. Infections caused by bacteria that have acquired resistance to an antibiotic can no longer be treated with that antibiotic. Consequently, identifying the type of pathogenic bacteria causing an infection may not always be sufficient to determine which antibiotics will be effective treatments. That is why all isolates must be tested to determine which antibiotics are effective before definitive treatment can be prescribed.
Activity 6: Comparing intrinsic and acquired resistance
Look at the following statements in the table. Decide whether they are about intrinsic or acquired resistance or both and type your answer into the right-hand column.
Answer
The answers are as follows:
| Statement | Intrinsic resistance, acquired resistance or both? |
|---|---|
| Mechanism only present in a subpopulation of bacteria of a given type | Acquired resistance |
| Can be identified if the bacterial type is known | Intrinsic resistance |
| Normal for bacteria of that type | Intrinsic resistance |
| Limits treatment options | Both |
| Mechanism present in all bacteria of a given type | Intrinsic resistance |
| Occurs as a result of genetic mutation or horizontal gene transfer | Acquired resistance |
2.2.3 Multidrug resistance
Bacteria can acquire multiple resistance mechanisms, making them resistant to multiple antibiotics. This is known as multidrug resistance (MDR). MDR bacteria are often known as ‘superbugs’ (although it is important to note that being resistant does not make bacteria more pathogenic, just that it will be harder to treat) and they are a major concern because they severely limit available treatment options.
Multidrug resistance is increasing globally, both in human healthcare and in farm animals. The issues in human healthcare are obvious, but multi-drug resistance in the food chain could have a severe impact on our ability to treat animals, as well as consumers of affected meat, in the future.
Activity 7: Acquiring multidrug resistance
In this activity you are asked to read a short article and answer some questions about it. One article is relevant to animal health and the other to human health; select the article that is most appropriate for your job and your interests.
Animal health
Read the Medical News Today article ‘Antibiotic resistance in farm animals is rising fast’, which highlights that, although multidrug resistance is rare, it is increasing and could cause significant problems for treatment of both animals and humans in the future.
While you read the article, note down the answers to the following questions.
- What percentage of antibiotics used in human treatment are routinely used in meat production?
Answer
73% are used in animals raised for food.
- Which countries have the largest number of multidrug resistance cases among farm animals?
Answer
India and north-east China have the most cases. Kenya, Uruguay and Brazil follow close behind.
- How much has the use of antibiotics in the last ten years affected resistance of bacteria found in samples from cattle, chickens and pigs?
Answer
The quantity of antimicrobials that bacteria found in cattle are resistant to has doubled. For chickens and pigs, the resistance is almost three times higher.
Human health
Read the BBC article ‘Bug resistant to all antibiotics kills woman’, which highlights that, although multidrug resistance is rare, it can have a devastating impact.
While you read the article, note down the answers to the following questions.
- Which bacterium caused the patient’s infection?
Answer
The patient’s infection was caused by the Gram-negative bacterium Klebsiella pneumoniae.
- How many antibiotics was the infection resistant to?
Answer
It was resistant to 26 different antibiotics, including the ‘drug of last resort’: colistin.
- Is resistance to all antibiotics a common occurrence?
Answer
No. Infections that are resistant to all antibiotics are uncommon.
3 Why are so many bacteria resistant to antibiotics?
Acquired resistance can result from mutations in chromosomal DNA or through
3.1 How do mutations lead to resistance?
A bacterium can develop antibiotic resistance through
How does altering the sequence of a bacterium’s DNA result in antibiotic resistance? The answer lies in how genetic information, encoded by DNA, is converted into proteins that are required for the structure and function of bacteria.
Optional activity: What do genes do?
If you are unfamiliar with the terms ‘DNA’, ‘RNA’, ‘base pair’, ‘gene’, ‘amino acid’ or ‘protein’, you may want to try our free OpenLearn course What do genes do? before you begin the following sections.
3.1.1 From genetic information to protein function
Almost every process in a cell requires proteins. As you saw previously, antibiotics often exert their bactericidal and bacteriostatic effects by binding to proteins that are crucial to the structure or function of the bacterial cell.
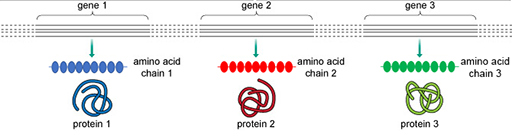
The function of a protein is largely determined by its structure. Proteins are composed of building blocks called amino acids. The sequence of these amino acids determines the structure of a protein. The
In 1958, Francis Crick, who helped discover the structure of DNA, proposed an explanation regarding how genetic information, encoded in DNA, can be converted into a functional product, a protein. In the first phase, a strand of DNA is used as a template to produce a complementary strand of RNA (ribonucleic acid). This process is called transcription. The RNA strand is then translated into a sequence of amino acids to form a new protein.
3.1.2 Genetic mutations and protein structure
As you already know, changes in the structure of bacterial proteins can result in antibiotic resistance.
Can you think of a specific example of how changing protein structure could lead to antibiotic resistance?
Structural changes to an antibiotic target protein could prevent that antibiotic from binding. This would make the target insensitive to the antibiotic and bacteria containing this protein would be resistant to the effects of that antibiotic. For example, linezolid exerts its antibiotic effects by binding to ribosomes and preventing the initiation of protein synthesis. Structural changes to the ribosome can prevent the binding of linezolid. Consequently, protein synthesis initiation is no longer blocked in the presence of linezolid and these resistant bacteria can grow.
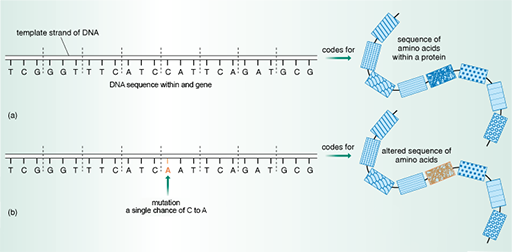
Remember that the amino acid sequence, and therefore the structure of a protein, is encoded in the DNA sequence of a gene. Small changes, or mutations, in the DNA sequence within a gene can alter the amino acid sequence of the protein it encodes (Figure 20).
Some changes in the bacterium’s DNA sequence have no effect on the amino acid sequence. Sometimes, however, even though the change in DNA is very small – perhaps a single point mutation – there can be a major effect on the amino acid sequence and, therefore, the structure of proteins that are targeted by antibiotics. As you have seen, these changes in the structure of proteins targeted by antibiotics can have important consequences for their function.
3.1.3 Transmission of mutations by vertical gene transfer
Activity 8: Exploring vertical transmission
Apply what you have learned to complete the following sentences. Select the appropriate word from the list.
a.
identical
b.
similar
c.
different
The correct answer is a.
Answer
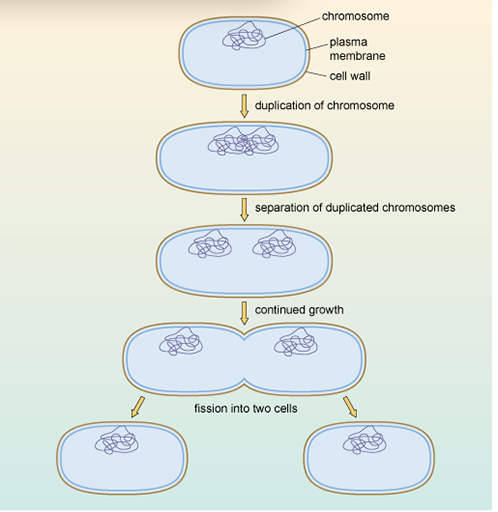
During binary fission, the genetic material (DNA) is copied so that each new daughter cell inherits an exact copy of the parent cell's DNA.
a.
sometimes
b.
never
c.
always
The correct answer is c.
Answer
During binary fission, the genetic material (DNA) is copied, so that each new daughter cell inherits an exact copy of the parent cell’s DNA. When the parent DNA is copied during binary fission, any genetic mutations will also be copied, and consequently inherited, by both of the daughter cells.
a.
sometimes
b.
always
c.
never
The correct answer is b.
Answer
During binary fission, the genetic material (DNA) is copied, so that each new daughter cell inherits an exact copy of the parent cell’s DNA. When the parent DNA is copied during binary fission, any genetic mutations will also be copied, and consequently inherited, by both of the daughter cells. If these genetic mutations give rise to antibiotic resistance in the parent bacteria, they will also result in antibiotic resistance in both of the daughters.
Vertical gene transfer is only one of the ways in which bacteria can spread antibiotic resistance genes. In the next section you will look at another – horizontal transfer.
3.2 Horizontal transfer
Horizontal gene transfer, or horizontal transfer, is the primary mechanism of spread of antibiotic resistance that allows bacteria to spread antibiotic resistance genes rapidly between different bacteria, and can sometimes include transfer to different bacterial species. As you should be starting to appreciate, the acquisition of antibiotic resistance by new bacterial types is particularly concerning because it can result in multidrug-resistant bacterial strains such as MRSA.
3.2.1 Plasmids
You have seen how chromosomal DNA can be copied and transmitted to the next generation via vertical gene transfer. Unlike humans and other animals, bacteria contain additional, non-chromosomal DNA which can be replicated independently of the genomic chromosomal DNA. These non-chromosomal genetic elements are called plasmids.
A
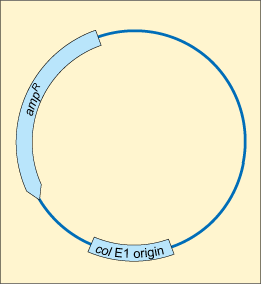
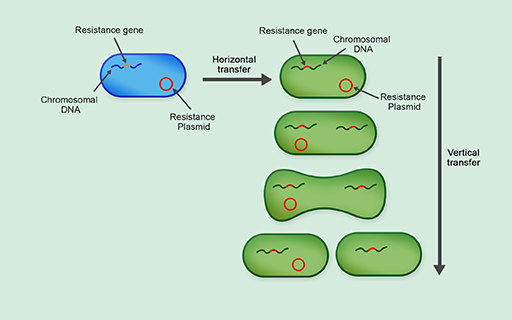
Plasmids are often transferred by horizontal gene transfer. This is the process of transferring genetic information between two unrelated cells. In contrast to vertical gene transmission, where plasmids are transferred from parents to daughter cells, it does not require binary fission and can occur between bacteria of the same generation, and even between bacteria of different species (Figure 24).
Can you suggest why horizontal gene transfer is the primary mechanism of spreading antibiotic resistance?
Horizontal gene transfer allows plasmids carrying antibiotic resistance genes to spread rapidly between different types of bacteria. This means that species of bacteria that are susceptible to a given antibiotic rapidly acquire resistance genes, making them resistant to treatment with that antibiotic.
There are three mechanisms of horizontal gene transfer:
- conjugation
- transformation
- transduction.
You will now look at each mechanism in more detail.
3.2.2 Conjugation
In the process of conjugation, plasmids are transferred between two contacting bacteria via a hollow tube or pilus (Figure 25).
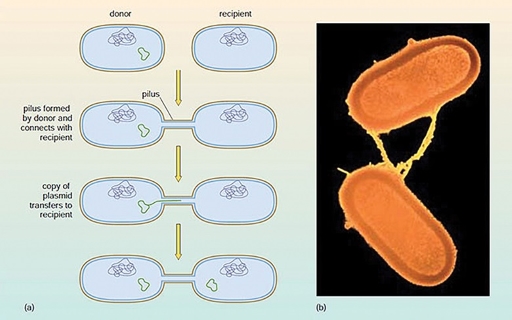
Since antibiotic resistance genes are often located on plasmids, conjugation can result in the transfer of antibiotic resistance from one bacterium to another.
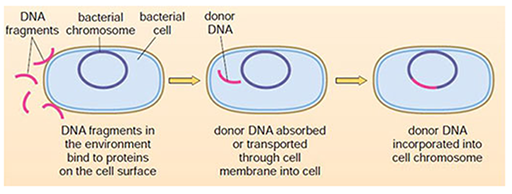
3.2.3 Transformation
In contrast to conjugation, the process of transformation allows bacteria to take up DNA from their environment (for example, from a lysed bacterium) across the cell wall. This DNA can then be incorporated into the genome of the bacterium (Figure 26).
Transformation occurs naturally between some bacteria, such as Streptococcus pneumoniae and Haemophilus influenza. When antibiotic resistance genes in the environment are transformed into a new bacterial type, they can be incorporated into that bacterium’s genome. They are then transmitted to the next generation by binary fission, establishing a newly resistant population of bacteria. Unincorporated plasmids are also usually (but not always) passed to the next generation.
3.2.4 Transduction
The final mechanism of horizontal gene transfer you will look at is
Viruses that infect bacteria are called bacteriophages. When a
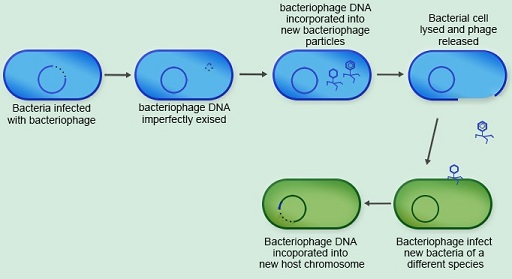
Activity 9: Comparing horizontal transfer mechanisms
Use the drop-down options to complete the five sentences below.
3.3 Evolving resistance to antibiotics
Within a mixed bacterial population (for example, in the gastrointestinal system), some bacteria may be susceptible to antibiotic treatment while others are intrinsically resistant, or may have acquired resistance to antibiotics, via either genetic mutation or horizontal gene transfer. In the presence of antibiotics, the resistant bacteria have a survival advantage over the susceptible bacteria and are more likely to survive and reproduce. Because bacteria reproduce so quickly, resistant bacteria can quickly dominate the population (Figure 28).
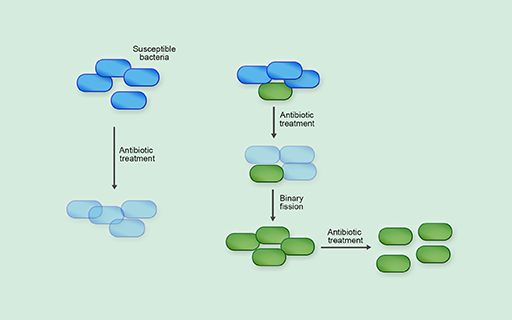
Of course, changes to the bacteria’s environment made by us can affect the evolution of antibiotic resistance.
3.3.1 Case study – antibiotics relevant to your work
Throughout this module you have studied the mechanism of action of several classes of antibiotics. In Activities 10–12 you will use an online database and apply what you have learnt to find out about the mechanism of action of an antibiotic related to your work.
In the following activity you will use the online Comprehensive Antibiotic Resistance Database (CARD) to find out some basic information about the mechanism of action of antibiotics relevant to your work.
Activity 10: Identifying antibiotics relevant to your work
You are asked to read a short paragraph and answer some questions about it. One paragraph is relevant to human health and the other to animal health. Select the paragraph that is most appropriate for your job and your interests. Identify three antibiotics that are relevant to your work.
Animal health
Mastitis is extremely common in dairy cattle, and is very costly for farmers. It is usually an infection of the mammary gland that can be caused by physical injury, but the most common cause is environmental contamination. Antibiotics available to treat bacterial mastitis include sulphonamides (an example of which is sulphanilamide), penicillin and streptomycin.
You can either use the mastitis example above or choose another disease from your work and the related antibiotics used to treat it. You may wish to talk to a colleague or supervisor to find out what diseases are commonly encountered in your workplace for ideas.
Use the space below to make a note of any antibiotics used in your work.
Activity 11: Accessing CARD
Now access the Comprehensive Antibiotic Resistance Database (CARD).
Type the name of one of the antibiotics that you identified in Activity 10 into the search box on the top right of the CARD website (highlighted in Figure 29).
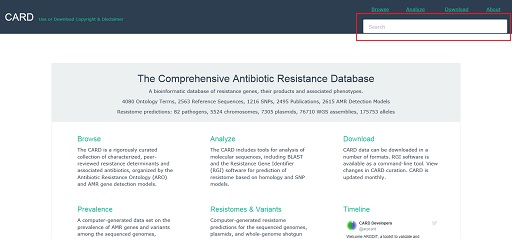
Select the antibiotic name from the resulting drop-down list (see Figure 30).
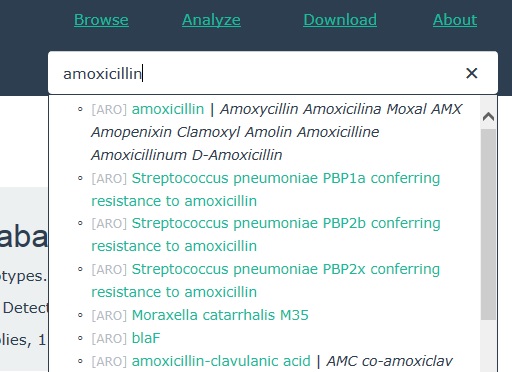
This should open a webpage containing information about the antibiotic. An example is shown in Figure 31.
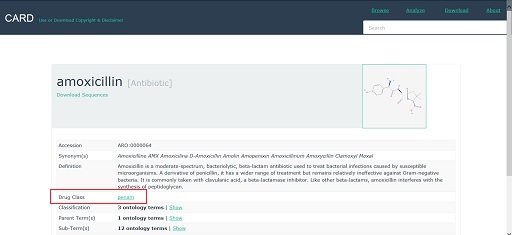
Explore the entry for the antibiotic that you have selected. Note down in the space below any information you can find about:
- the drug class that the antibiotic belongs to
- whether its action is bacteriostatic or bactericidal
- whether it is active against Gram-negative or Gram-positive bacteria, or both
- whether it is broad- or narrow-spectrum
- the bacterial process that it interferes with.
Be aware that some information about the antibiotic may be given on the drug class page. To access this information you will need to click on the name of the drug class to open the link (highlighted in Figure 31).
Do not worry if you cannot find all of this information in the CARD database. You could also try searching online or asking your colleagues for information. Now repeat the process for two other antibiotics used to treat your chosen disease or pathogen.
Activity 12: Showing what you have learnt
Write a short summary (of four or five sentences) about what you have found out. Include the name of the antibiotic and why you have selected it. To help you, an example of a forum post is given below:
Animal health example
Bacterial diarrhoea, or scour, in piglets can be treated using cephalexin as long as the bacterial pathogen is susceptible to it. In my workplace we carry out regular tests to determine whether diarrhoeal samples from pigs from the local farms are resistant to cephalexin. Cephalexin is a cephalosporin antibiotic. It is a broad-spectrum, bactericidal antibiotic which acts by inhibiting bacterial cell wall synthesis. It is active against both Gram-positive and Gram-negative bacteria.
Human health example
In my workplace we carry out regular tests to determine whether diarrhoeal samples from the local community are resistant to cephalexin. Cephalexin is a cephalosporin antibiotic. It is a broad-spectrum, bactericidal antibiotic which acts by inhibiting bacterial cell wall synthesis. It is active against both Gram-positive and Gram-negative bacteria.
Once you have written your three summaries, spend a few minutes comparing the different antibiotics and how they act. Are they from different classes, or the same? Do they have the same mechanism of action or different? What else is similar or different about them?
4 End-of-module quiz
Well done – you have reached the end of this module and can now do the quiz to test your learning.
This quiz is an opportunity for you to reflect on what you have learned rather than a test, and you can revisit it as many times as you like.
Open the quiz in a new tab or window by holding down ‘Ctrl’ (or ‘Cmd’ on a Mac) when you click on the link.
5 Summary
In this module you looked at the main modes of antibiotic action and learned why these drugs work selectively against bacteria without causing the same level of harm to the host. You have also learned that bacteria can be intrinsically resistant to antibiotics or acquire antibiotic resistance that can rapidly spread, or evolve, in a population. You should now be able to explain how genetic mutations cause acquired antibiotic resistance and how these mutations can be inherited through binary fission.
You have also seen how horizontal gene transfer has an important role in transmitting antibiotic resistance to different bacterial types. Having seen how antibiotic resistance evolves to protect bacteria, you can now begin to speculate on how our use of antibiotics contributes to the rise of antibiotic resistance.
You should now be able to:
- explain how antibiotics work against bacterial pathogens
- state what is meant by the term ‘antibiotic resistance’
- explain that antibiotic resistance is a natural phenomenon that protects bacteria from hostile environments
- explain how resistance mechanisms arise, and how resistance can be passed on during bacterial replication
- describe the horizontal gene transfer mechanisms that allow antibiotic resistance to be transferred between bacteria
- apply scientific terminology when explaining how antibiotic resistance relates to your current work.
Now that you have completed this module, consider the following questions:
- What is the single most important lesson that you have taken away from this module?
- How relevant is it to your work?
- Can you suggest ways in which this new knowledge can benefit your practice?
When you have reflected on these, go to your reflective blog and note down your thoughts.
Activity 11: Reflecting on your progress
Do you remember at the beginning of this module you were asked to take a moment to think about these learning outcomes and how confident you felt about your knowledge and skills in these areas?
Now that you have completed this module, take some time to reflect on your progress and use the interactive tool to rate your confidence in these areas using the following scale:
- 5 Very confident
- 4 Confident
- 3 Neither confident or not confident
- 2 Not very confident
- 1 Not at all confident
Try to use the full range of ratings shown above to rate yourself:
Reflect on your progress by comparing your answers here to those you wrote at the beginning of the module. Have your responses changed?
Remember in Activity 2 where you were asked to construct your own glossary of terms? Has this helped to expand your understanding of your scientific knowledge relating to your own working environment?
When you have reflected on your answers and your progress on this module, go to your reflective blog and note down your thoughts.
6 Your experience of this module
Now that you have completed this module, take a few moments to reflect on your experience of working through it. Please complete a survey to tell us about your reflections. Your responses will allow us to gauge how useful you have found this module and how effectively you have engaged with the content. We will also use your feedback on this pathway to better inform the design of future online experiences for our learners.
Many thanks for your help.
References
Acknowledgements
This free course was written by Kay Saunders and Rachel McMullan, with contributions from Ben Amos, Claire Gordon, Natalie Moyen and Hilary MacQueen.
Except for third party materials and otherwise stated (see terms and conditions), this content is made available under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 Licence.
The material acknowledged below is Proprietary and used under licence (not subject to Creative Commons Licence). Grateful acknowledgement is made to the following sources for permission to reproduce material in this free course:
Images
Course image: Dr_Microbe/iStock/Getty Images Plus.
Figures 1–6, 8a, 9–13, 15–21, 23–25a and 26–28: The Open University.
Figure 7: Male, D. et al., Immunology, 7th edn, Elsevier.
Figure 8b: CNRI/Science Photo Library.
Figure 14: adapted (text correction) from Drakosam, ‘Mechanism of daptomycin action’, https://commons.wikimedia.org/ wiki/ File:Mechanism_of_Daptomycin_Action.jpg. This file is licensed under a Creative Commons Attribution-ShareAlike 4.0 International (CC BY-SA 4.0) licence, https://creativecommons.org/ licenses/ by-sa/ 4.0/ deed.en.
Figure 22: Nick Kim/CartoonStock.
Figure 25b: Dr L. Caro/Science Photo Library.
Figures 29–31: Jia et al. (2017) ‘CARD 2017: expansion and model-centric curation of the Comprehensive Antibiotic Resistance Database’ Nucleic Acids Research, 45, D566–73 [PMID 27789705]. © McMaster University.
Videos
Video 1: courtesy e-bug (https://www.e-bug.eu/).
Video 2: BBC, ‘Michael Mosley vs The Superbugs’, TX 17 May 2017.
Video 3: The Open University.
Every effort has been made to contact copyright owners. If any have been inadvertently overlooked, the publishers will be pleased to make the necessary arrangements at the first opportunity.