Use 'Print preview' to check the number of pages and printer settings.
Print functionality varies between browsers.
Printable page generated Wednesday, 15 May 2024, 4:29 AM
Testing for mechanisms of resistance
Introduction
In this module you will learn about phenotypic and genotypic laboratory techniques that are used to test for common mechanisms of antimicrobial resistance (AMR). World Health Organization (
You will start by looking at resistance patterns of global concern in human and animal health, and the importance of knowing the underlying resistance mechanism involved. You will further learn how data derived from screening for specific resistance mechanisms and confirmatory testing, differs from
Basic knowledge of concepts such as the emergence and spread of ‘
By the end of this module you should be able to:
- give examples of the resistance patterns/resistant organisms causing global concern
- give examples of the resistance patterns/resistant organisms encountered in your work
- understand the difference between screening and confirmatory testing
- describe some of the phenotypic methods commonly used for screening and confirmation of resistance mechanisms
- understand how genotypic methods can be used, and the advantages/disadvantages of genotypic versus phenotypic methods
- understand how more data from detailed testing for resistance mechanisms/genes contributes to AMR surveillance
- apply your knowledge of these laboratory tests to interpret data relevant to your work
- know the importance of procedures designed to ensure the quality of these laboratory tests in your workplace.
Activity 1 Assessing your skills and knowledge
Before you begin this module, think about your current level of knowledge and skills in the areas covered in this module. You will have an opportunity to repeat this activity when you have completed the module. Do not worry if you lack confidence in some areas, they may be knowledge areas and skills that you are hoping to develop by studying this module. For areas where you feel fully confident, it is always a good idea to refresh and update knowledge and skills.
Use the interactive tool to rate your confidence in these areas using the following scale:
- 5 Very confident
- 4 Confident
- 3 Neither confident nor not confident
- 2 Not very confident
- 1 Not at all confident
This is for you to reflect on your own knowledge and skills you already have.
1 Resistance problems of global concern
1.1 Priority human and animal pathogens
Bacteria isolated from clinical sources and animals may have acquired resistance to one or more antimicrobials. A minority of these are both
Of the eight WHO-listed human pathogens which are the focus of the Global Antimicrobial Resistance Surveillance System (
In contrast, different bacteria are the main focus of active AMR surveillance in healthy animals, of which some are commensals and some zoonotic pathogens (EFSA, n.d.; FAO, n.d.). In some countries, such as France, Denmark and the USA, bacterial pathogens from certain animal species are included in routine AMR surveillance, for example, livestock-associated MRSA in pigs and Salmonella in cattle (CDC, 2019; DVFA, n.d.). AMR data on these bacterial pathogens are generated from routine laboratory investigations when animals display clinical disease and from controls at abattoirs on healthy animals.
-
Can you recall the names of any GLASS pathogens? [For clinicians]
-
The eight key bacterial pathogens identified by WHO as the focus of GLASS AMR surveillance programmes are:
- Staphylococcus aureus
- Streptococcus pneumoniae
- Acinetobacter species
- Escherichia coli
- Klebsiella pneumoniae
- Neisseria gonorrhoeae
- Salmonella species
- Shigella species
-
Can you recall the names of any commensal or pathogenic bacterial flora in animals which are important causes of zoonotic infection and/or introduction of resistant bacteria via the human food chain? [For veterinarians]
-
Organisms of concern include:
- two serotypes of Salmonella, Enteritidis and Typhimurium
- Campylobacter species
- Enterococcus species, especially E. faecium and E. faecalis;
- E. coli
- K. pneumoniae
For background information see the EFSA Panel on Biological Hazards (2009) and WHO (2018).
1.2 WHO priority human pathogens for antimicrobial R&D
The WHO (2017a) has also developed a different list of specific organism-resistance combinations which have the most urgent need for research and development (R&D) into new treatment approaches. Nine of the twelve organisms listed are
Priority 1: CRITICAL
- Acinetobacter baumannii – carbapenem-resistant
- Pseudomonas aeruginosa – carbapenem-resistant
- Enterobacterales – carbapenem-resistant, ESBL-producing.
Priority 2: HIGH
- Enterococcus faecium – vancomycin-resistant (VRE)
- Staphylococcus aureus, – methicillin-resistant (
MRSA ), vancomycin-intermediate and vancomycin-resistant (VISA ,VRSA ) - Helicobacter pylori – clarithromycin-resistant
- Campylobacter species – fluoroquinolone-resistant
- Salmonella species – fluoroquinolone-resistant
- Neisseria gonorrhoeae – cephalosporin-resistant, fluoroquinolone-resistant
Priority 3: MEDIUM
- Streptococcus pneumoniae – penicillin-non-susceptible
- Haemophilus influenzae – ampicillin-resistant
- Shigella species – fluoroquinolone-resistant
-
Why do you think these organisms were chosen?
-
Selection criteria for the WHO priority R&D pathogen list (WHO, 2017b) were:
- the severity of the infections caused
- whether treatment of the infection requires a long hospital stay
- how frequently the organism is resistant to existing antimicrobials when people in communities catch them
- how easily the organism spreads between animals, from animals to humans, and from person to person
- whether the infection can be prevented, for example through good hygiene and/or vaccination
- whether many treatment options remain
- whether new antimicrobials to treat them are already in the R&D pipeline.
Activity 2 GLASS versus priority R&D WHO lists
Compare the organisms featured on the WHO GLASS and the R&D lists and make notes of any similarities and differences. What are the reasons for any differences between the two lists?
Discussion
You will have noticed that there is some overlap between the lists but also some striking differences.
At a basic level, the GLASS list applies to species and looks at a range of resistance mechanisms rather than specifying single species-AMR combinations. This is because the lists serve different functions.
The WHO priority R&D list is about the need for new antimicrobials. It addresses the most concerning resistant organisms known currently. Some of these, like fluoroquinolone-resistant Salmonellae are mainly found in the community while others, for example VRE E. faecium, are mostly found in tertiary care settings.
In contrast, GLASS monitors a wider range of agents as its purpose is to track the development and spread of AMR overall including new patterns of resistance important in human medicine. GLASS is concerned with AMS and the organisms on the list are common and widespread. They can be cultured in most hospital microbiology laboratories, monitored and potentially controlled by good infection control and AMS.
Some rarely encountered organisms like VRSA are commonly tested for in clinical laboratories, while organisms like H. pylori, which are very widespread in the population, are not. Attempts to culture H. pylori are rare in routine clinical laboratories, so it would not be practical to include these in the GLASS surveillance.
Activity 3 WHO priority R&D list in context of One Health
Which pathogen-resistance combinations in the WHO priority R&D list are also important from a
Discussion
The WHO R&D priority list includes several pathogens of One Health importance which are found in livestock:
- Enterobacterales – carbapenem-resistant, ESBL-producing
- Enterococcus faecium – vancomycin-resistant (VRE)
- Staphylococcus aureus – methicillin-resistant (MRSA), vancomycin-intermediate and resistant (VISA, VRSA); the latter are unlikely to be detected in routine enteric samples but can be cultured from samples of animal origin
- Campylobacter species – fluoroquinolone-resistant
- Salmonella species – fluoroquinolone-resistant.
1.3 Resistance patterns in your workplace
Activity 4 Resistance patterns in your workplace [For clinicians]
Think about the organisms your laboratory identifies routinely. Referring to the GLASS and WHO priority R&D lists, what are the most important organism-resistance patterns that you see in your workplace? Are any combinations associated with outbreaks, for example in Critical Care? Do you see any patterns regularly which are not on the lists? If you routinely perform AST, would your laboratory currently be able to detect the resistance profiles of concern? Make notes and then compare with the example answer.
Discussion
It is likely that you encounter MRSA and ESBL-producers as these are very common, as well as carbapenemase producers if you live in a part of the world where these are established. If you have a Critical Care Unit in your hospital you are likely to have seen outbreaks with some of the MDR Gram negatives.
Activity 4 Resistance patterns in your workplace [For veterinarians]
Think about the organisms your laboratory identifies routinely. Have you had experience in detecting any of the organisms from the WHO R&D priority list in your laboratory? If you routinely perform AST, would your laboratory currently be able to detect the resistance profiles of concern? Make notes and then compare with the example answer.
Discussion
You are likely to be identifying the coliforms and enteric pathogens. Resistance is surprisingly common if you test for it, both in animals which have been treated with antimicrobials and in those which have not. The strains are readily transmissible and can be spread from other animals via the environment or from humans, hence the need to test for resistance.
Most of the GLASS and WHO priority R&D pathogens, including those of One Health importance, can be cultured easily; some have specific growth requirements (see the Isolating and identifying bacteria module). Some of the resistance patterns can be identified by routine AST; others, for instance ESBL production or VRE, may need additional testing to identify and confirm their presence. You will find out more about these methods later in the module.
2 Antimicrobial resistance
2.1 Resistance mechanisms
The resistance mechanisms that bacteria have evolved to counter the action of antimicrobials are sophisticated. In the Introducing antimicrobial resistance module you learned about the main mechanisms of antimicrobial resistance. See if you can remember them in Activity 5.
Activity 5 Antimicrobial resistance mechanisms
Part 1
Figure 2 shows an overview of the main mechanisms of antimicrobial resistance. Use the drop-down list to match the mechanism to the appropriate part of the diagram.
Discussion
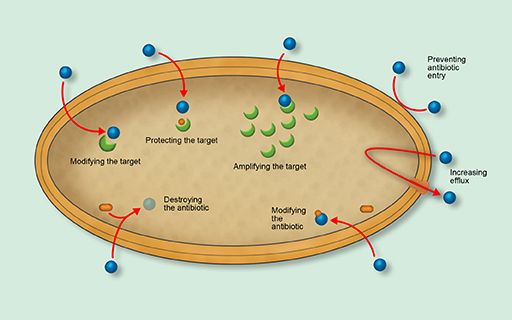
- Preventing antimicrobial entry
- Increasing efflux
- Modifying the antimicrobial
- Destroying the antimicrobial
- Modifying the target
- Protecting the target
- Amplifying the target.
Unlike hydrolysing enzymes which in general provide high levels of resistance, some mechanisms only produce low-level resistance on their own. However, different mechanisms can combine to increase the level of resistance provided. For example, porin loss combined with an over-expression of efflux pumps can lead to carbapenem resistance which is hard to distinguish on AST from the hydrolysing activity of a carbapenemase, and which has the same effect clinically in reducing the efficacy of treatment.
The β-lactam-hydrolysing enzymes are interesting in that they can sometimes be blocked by using a second substance along with the antimicrobial. This enzyme blocker is called an
2.2 Spread of antimicrobial resistance in bacteria
Some species of bacteria have
Activity 6 Plasmids
From your study of the Introducing antimicrobial resistance module what can you remember about the role plasmids play in the spread of antimicrobial resistance? Make notes in the text box provided.
Discussion
- Plasmids are mobile genetic elements easily transmitted between bacteria, including between different strains and species.
- Plasmids can readily acquire multiple resistance genes which can quickly lead to MDR transmissible strains, making them important in the spread of resistance.
- Plasmid acquisition, unlike the accrual of multiple mutations, confers immediate high-level resistance to one or often, multiple, antimicrobials and threatens the ability to treat severe infections.
- Antimicrobial use provides selection pressure on bacteria to retain their plasmids.
Resistance can be shuffled between chromosomal and plasmid DNA on small mobile genetic elements, facilitating recombination and promoting the rapid evolution of new resistant forms. Resistance genes can remain on the plasmid or be integrated into the chromosome; chromosomal mutations can also find their way onto plasmids. This, coupled with
The current greatest AMR threat to modern medicine is from plasmids with a carbapenemase gene plus resistance to multiple antimicrobials. In any of the three organisms in the WHO ‘critical’ R&D category, carbapenem-resistance leads to infections that can only be treated with combinations of drugs or toxic ‘third line’ antimicrobials, for example, colistin. These plasmids can lead to untreatable infections, hence the high priority placed on these pathogen-AMR combinations.
3 Pathogen-AMR combinations
In this section we will explore in more detail some of the specific pathogen resistance patterns in human and animal health.
3.1 Gram-negative pathogens
Gram-negative pathogens are a significant cause of morbidity both in community-acquired infection such as enteric fever and urinary tract infection (
In animal health, Gram-negatives are found as gut commensals and as pathogens. They are an important source of resistance genes getting into the human food chain, especially as it is hard to totally prevent contamination of meat with faecal material at slaughter.

Production of β-lactamases and carbapenemases are the biggest problem with infections caused by Gram-negative organisms owing to the relative lack of antimicrobials available outside these classes to treat them. Quinolone resistance is also a problem.
3.1.1 β-lactam resistance and ESBLs
The resistance genes for β-lactamases are located either on the bacterial chromosome or on plasmids, leading to both intrinsic and acquired resistance. The genes have mutated and evolved so that each one represents a ‘family’ of related genes. Two separate systems are used to classify β-lactamases: a structural scheme based on primary structure (Ambler, 1980; Hall and Barlow, 2005); and a functional scheme based on chemical reactivity and DNA sequence (Bush and Jacoby, 2005). The Ambler classification is based on classes A-D (Table 1), whereas the Bush‒Jacoby has numbered groups. This may seem complex but fortunately only a few subdivisions of the main classes of β-lactamases are widespread enough to mention specifically here in the context of surveillance.
| Ambler DNA sequence classification of β-lactamases | ||
|---|---|---|
| Class | Active site | Selected examples |
| A | Serine | Staphylococcal penicillinase Broad-spectrum penicillinases, TEM and SHV ESBLs e.g. CTX-TM, TEM-3 and SHV-2 Serine carbapenemases, e.g. KPC. |
| B | Zinc | Metallo-carbapenemases, e.g. IMP, NDM, VIM |
| C | Serine | Chromosomal cephalosporinases (some genes may be on plasmids), e.g. major families of plasmid-carried genes for AmpC such as CMY, LAT, FOX. |
| D | Serine | Oxacillinases - broad-spectrum not inhibited by clavulanate or tazobactam, e.g. OXA-1 and OXA-48 carbapenemase. |
ESBL-producers among Enterobacterales are significant as they cannot be treated with the usual ‘
ESBL-producers, mainly E. coli and Klebsiella, are also widely distributed in poultry, including those which have had little direct exposure to antimicrobials themselves. They are readily transmitted to humans via food consumption, for example by handling uncooked meat or consuming vegetables fertilised with animal manure; they can also be transferred in the other direction from humans to livestock (Subramanya et al., 2020; Uttapoln et al., 2019).
Cephalosporin hydrolysing enzymes of clinical and veterinary importance are shown in Table 2.
| Group | Notes |
| CTX-M | ESBL: Found in E.coli and Klebsiella species; widespread and the predominant ESBL variants in many countries. The gene originated in an environmental organism, Kluyvera.(Class A) |
| TEM and SHV | ESBL: Increasingly diverse with some being able to break down carbapenems as well as cephalosporins. They have been present in clinical isolates for many years which explains why the group is so diverse. (Class A). |
| AmpC | Common in Enterobacterales. AmpC is not affected by inhibitors. Many species have the ampC gene on their chromosome but as this resistance is intrinsic it is not important for surveillance. The AmpC enzyme can, however, mask the presence of transmissible genes. Acquired plasmid-mediated ampC is of importance in a public health context, because unlike intrinsic ampC, it can be passed on to other bacteria. (Class C) |
Table 2 Cephalosporin hydrolysing enzymes important in human and animal health
3.1.2 Carbapenem resistance and carbapenemases
The greatest current concern for human health is from carbapenem resistance. For example, Acinetobacter, Klebsiella and Pseudomonas aeruginosa are important causes of outbreaks in tertiary and critical care. MDR- and pan-resistant strains are a problem in many critical care units. They are frequently resistant not just to carbapenems but also to all other available antimicrobials. Once they have colonised the environment in a unit, they are very hard to get rid of.
Carbapenem-resistant Enterobacterales (CPE), predominantly Klebsiella and E. coli, have been found in wild animals, livestock and pets, with evidence of direct transfer to humans (Kock et al., 2018).
The main classes of carbapenemases of clinical and veterinary importance are shown in Table 3.
| Carbapenemase class | Notes |
| KPC | Found worldwide, particularly associated with Klebsiella |
| VIM | Metallo-β-lactamases mainly found in Pseudomonas |
| IMP | Metallo-β-lactamases found in Pseudomonas, Acinetobacter and Enterobacterales |
| NDM | Metallo-β-lactamases most common at present in India and the Middle East in Enterobacterales but spread to many species. |
| OXA-48 | Most common carbapenemases in many parts of the world and evolving and spreading rapidly. Most important in K. pneumoniae and E. coli but also found in Pseudomonas where they are harder to detect. They can be missed if carbapenemase screening is not done correctly. |
Table 3 Carbapenemases important in human and animal health (abbreviations: KPC – Klebsiella pneumoniae carbapenemase; VIM – Verona integron-encoded metallo-beta-lactamase; IMP – Imipenemase metallo-beta-lactamase; NDM – New Delhi metallo-β-lactamase; OXA-48 – Oxacillin-48 carbapenemase)
3.1.3 Fluoroquinolone resistance
Fluoroquinolone antimicrobials are available orally including for severe infections. They can be used for intracellular organisms – in particular, S. typhi against which many other antimicrobials are not effective as they are unable to enter the mammalian cell – and are a very useful class of antimicrobials. However, fluoroquinolone resistance among Gram-negative organisms is widespread and two pathogen-AMR combinations are listed in the WHO ‘high’ category for R&D priority: fluoroquinolone-resistant Salmonellae and Campylobacter species. Fluoroquinolone-resistant Shigella species are of ‘medium’ R&D priority. Resistance is mostly via modification of the antimicrobial target, as a result of one or more chromosomal mutations.
Both fluoroquinolone-resistant Campylobacter and Salmonella infections are major zoonoses and there are few antimicrobial treatment options. The infections can easily be acquired from food of animal origin if food hygiene is not strictly applied. In contrast, fluoroquinolone-resistant Shigella infections are not acquired from food animals and in general, only affect humans and other primates. Reliance is on good hygiene and sanitation to prevent spread, so they are widespread in contexts where this is difficult to achieve.
3.2 Gram-positive pathogens
In human health, two
3.2.1 β-lactam resistant S. aureus
Resistant strains of S. aureus pose a big threat to both human and animal health worldwide, with MRSA S. aureus resistant to β-lactam antimicrobials of particular importance.
MRSA is easily transmitted between patients and is a very important HCAI pathogen globally.
MRSA is also found increasingly in livestock (Livestock-associated MRSA,
Resistance occurs by two main mechanisms. The most common form is due to the presence of the mecA gene on a mobile genetic element carried by all MRSA strains. MecA encodes an altered
This type of resistance is different from staphylococcal β-lactamases. These enzymes only cause resistance to penicillins (pencillin, ampicillin and amoxicillin) and the organism remains susceptible to isoxazolyl-penicillins (flucloxacillin and cloxacillin), and first generation cephalosporins. As such, these basic β-lactamase producers do not pose a public health problem at the moment as they remain relatively easy to treat.
3.2.2 Vancomycin-resistant S.aureus
Vancomycin is important against S. aureus as it can be used for MRSA strains or when β-lactams cannot be used. VISA refers to organisms with intermediate resistance while hVISA refers to subpopulations of organisms with intermediate resistance in a single culture. VRSA organisms are fully resistant to vancomycin. VISA, hVISA and VRSA therefore refer to S.aureus strains with varying degrees of resistance to vancomycin and using a number of resistance mechanisms. They have been found in increasing numbers in recent years in isolates from both humans and livestock and are a potential future threat (Al-Amery et al., 2019; Cong et al., 2020; Shariati et al., 2020).
Of greatest concern is the presence of the VanA genes, originally found in
3.2.3 Vancomycin-resistant Enterococci
Enterococci are gastro-intestinal commensals of humans and other animals. They are moderately virulent, mainly in the context of healthcare i.e. they are largely a HCAI pathogen, although they can cause urinary tract infection in healthy individuals. They cause infections related to prosthetic materials such as intravascular devices, mechanical heart valves and joint implants, that can be extremely difficult to treat.
VRE strains of Enterococcus, most commonly Enterococcus faecium, cause outbreaks among cancer and critical care patients with infections that are hard to treat and require expensive antimicrobials.
E. faecium is intrinsically resistant to β-lactams, including amoxicillin which is active against other Enterococci such as E. faecalis, which means that the loss of vancomycin sensitivity is important. Other species exist that are intrinsically resistant to vancomycin e.g. E. gallinarum and E. casseliflavus, but these only cause sporadic infections in humans (CDC, 2010).
There are a number of resistant phenotypes and genes involved in VRE, the main ones of clinical concern being those with the vanA phenotype. This is mediated by the vanA
Enterococci of animal origin are important from the One Health perspective as a source of resistance genes entering the human food chain.
3.3 Miscellaneous pathogens of interest
3.3.1 Respiratory and central nervous system pathogens
High-level penicillin resistance in S. pneumoniae can lead to treatment failure in respiratory tract infections. Even low-level resistance is a problem for infections of the central nervous system, for example meningitis. This is because only small concentrations of drug can be achieved in the brain because of the presence of the blood-brain barrier (Linares et al., 2010).
Penicillin resistance in S. pneumoniae is acquired differently to the resistance mechanisms described so far. It is generally the result of cumulative mutations and resistance acquisition leading to resistant strains, rather than by a single transmissible genetic element. An association has often been observed between rates of antimicrobial use and prevalence of resistance in this organism (Albrich et al., 2004; WHO, 2001).
-
Why is this organism in the ‘medium’ WHO priority R&D category and not in the ‘high’ category?
-
Although this organism is responsible for life-threatening pneumonia and meningitis, antimicrobial resistance is not as readily transmissible as for some of the other organisms. The resistant strains themselves are transmissible but infection can be controlled by vaccination, and as the strain is removed from circulation so too is its resistance. Most resistant strains remain susceptible to some antimicrobials so can still be treated. MDR S. pneumoniae is an emerging problem (Cillóniz et al., 2018) though, so it could be upgraded in a future list (WHO, 2001).
Haemophilus influenzae is a cause of vaccine-preventable invasive disease (HiB) in children, mainly causing respiratory tract infection or meningitis. Despite the ongoing success of the global vaccine programme, resistance to amoxicillin, in both HiB and non-typeable H. influenzae (NTHi) strains, remains a problem as amoxicillin would normally be the treatment of choice (Su et al., 2020; Van Eldere et al., 2014).
3.3.2 Sexually transmitted pathogens
MDR isolates of N. gonorrhoeae are increasing, with fluoroquinolone resistance a particular problem as these infections may not have any treatment options.
Resistance to third generation cephalosporins (ceftriaxone) is also starting to lead to strains which are almost impossible to treat, even with intramuscular injection of antimicrobials rather than oral ones (Costa-Lourenço et al., 2017).
4 Identifying resistance mechanisms
4.1 Why knowing the mechanism is important
-
Why might it be important to know what the resistance mechanism is for an isolate, rather than just whether it is susceptible to the antimicrobial you are testing?
-
Knowing the resistance mechanism is important from both a broader public health and a local infection control perspective. Some resistance mechanisms have more public health consequences than others, for example MRSA versus beta-lactamase producing S. aureus (see section 3.2.1 above) or transmissible
carbapenemases compared to efflux pump-mediated resistance, because of their ability to spread combined with high-level resistance to critically important antimicrobials.
Activity 7 What the resistance mechanism tells us
Imagine that a novel pathogen-antimicrobial combination has been found in your country or institution for the first time. What important questions from a public health or infection control perspective need answering? How might knowing the underlying resistance mechanism(s) of the isolate help answer these questions?
Make notes in the text box below before looking at the example answer.
Discussion
Important questions include:
- Is this mechanism new in this species?
- For example, a completely new type of plasmid-mediated colistin resistance gene (mcr-1) found in E. coli isolated from pigs was discovered in China in 2015. There are now nine variants of this resistance gene, mcr-1 to mcr-9 (Liu et al., 2016).
- Where did the resistance mechanism originate from? Has it jumped from another bacterial species?
- For example, although the mcr-1 gene’s origins are not yet known, others are, for example the CTX-M gene in Klebsiella came from a different Gram-negative organism (Kluyvera) originally.
- Is resistance transmissible? Is it on a plasmid?
- For example, the mcr-1 gene being plasmid-borne can easily transfer between different bacteria, and has now been reported in E. coli, Salmonella enterica, K. pneumoniae, and Enterobacter species. Since the discovery of the mcr-1 gene in China, it has now been reported in over 30 countries.
- Is it linked to other resistance genes? Many resistance genes are found on plasmids along with a wide range of other resistance genes leading to MDR organisms. If it has been found in animals, has it also spread to human populations?
- For example, the mcr-1 gene has moved from pig pathogens into pathogens found in people and the mcr plasmids are now circulating within populations of human pathogens.
- Can it still be contained? Or has it already spread too widely by the time it is detected?
- What infection control measures are advisable? Have similar pathogen-resistance combinations been controlled successfully in the past?
4.2 The approach used to determine the resistance mechanism
Once AST has confirmed that a bacterial isolate is resistant to antimicrobial(s), two approaches can be used, either separately or in combination, to identify the resistance mechanism involved ‒
Phenotypic tests focus on characteristics of the bacteria (i.e. phenotype) that can be readily observed or tested for in the laboratory. They use the results of these observations/tests to infer the resistance mechanism (and thus the gene) involved. In contrast, genotypic tests identify the gene responsible for the resistance directly.
You will learn more about these two approaches in the following sections.

5 Phenotypic tests
Phenotypic tests are carried out more frequently than genotypic tests as they are easier to perform and require less sophisticated/specialised equipment. They include the following, discussed in more detail below:
- use of indicator antimicrobials
- testing for inducible resistance
- synergy tests
- use of interpretative panels
- antigen detection tests
- colorimetric tests.
5.1 Use of indicator antimicrobials
Indicator antimicrobials can be used alongside normal AST. The principle of this test is to use the
In these instances, once you know that the organism is resistant to the indicator, you can then extrapolate that it will be resistant to every antimicrobial in that class. However, this only works in a few specific cases. For example, cefoxitin resistance in S. aureus implies mecA is present, so the Staphylococcus must be an MRSA (see Section 3.2.1).
Certain indicator antimicrobials can also be used to detect non-
5.2 Testing for inducible resistance
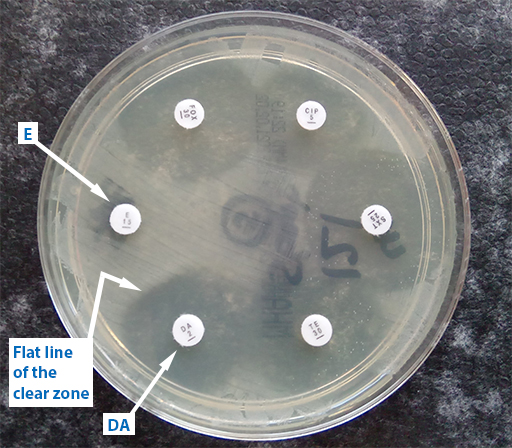
Some resistance mechanisms are expressed only when induced by the presence of another antimicrobial. These strains can become resistant during antibiotic treatment, so it is important to know if they are present. For certain pathogen-antimicrobial combinations, it is therefore possible to test for the resistance mechanism using a form of AST called the
The ‘D’ test can be used to detect inducible clindamycin (DA) resistance in S. aureus. In Figure 10, erythromycin (E) has diffused into the agar and interfered with the effect of the adjacent antibiotic, DA. The effect of E has been to induce the expression of DA in S.aureus organisms present in the agar and allow the bacteria to grow closer to the DA-impregnated disk. This is shown by flattening of the DA inhibition zone on the side next to the E disk.
5.3 Synergy tests
See Section 9.1.2 for examples of synergy testing.
5.4 Use of interpretative panels
Interpretative panels can be done as individual tests, but in practice they are mostly done using an automated system such as Vitek IITM, SensititreTM or PhoenixTM.
5.5 Antigen detection tests
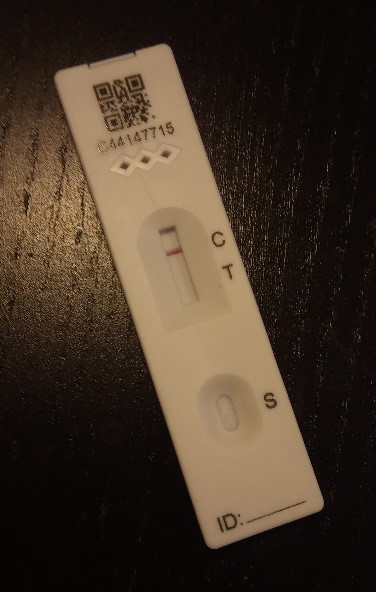
Antigen detection tests are mainly used to identify antibiotic hydrolysing enzymes, such as ESBLs and carbapenemases. The enzymes are detected by lateral flow tests to detect a specific enzyme.
The example test in Figure 12 is negative but has passed the control. A positive test would have a line by ‘T’ as well. Lateral flow tests are easy to perform though sometimes hard to interpret if the test line appears faintly.
5.6 Colorimetric tests
Colorimetric tests work on the principle that a substrate changes colour when broken down by an enzyme. This principle is used in chromogenic media (e.g. the commercially available CHROMagarTM agars developed for detection of carbapenemase producers, Acinetobacter, MRSA etc.). As well as having substrates for the enzymes, some of them are also selective, that is they inhibit the growth of susceptible organisms and species which are not of interest and only support those to be isolated. Some also change colour for a particular species so can be used to identify the species as well as the resistance mechanism.
Another example is nitrocefin – a chromogenic cephalosporin – which can be used to screen for β-lactamases (Video 1). Nitrocefin is broken down by all known β-lactamases leading to a colour change from colourless to red and so works as an indicator antimicrobial. Although it can be used to test whether a β-lactamase is present it will not discriminate between types of β-lactamase. It is routinely used to look for β-lactamase in N. gonorrhoeae and H. influenzae.
6 Genotypic tests
Genotypic tests are widely recognised as the best way to unambiguously prove that a specific resistance gene is present. They are also the only way of characterising suspected new resistance mechanisms.
Many genotypic tests are only done in research and reference laboratories as they all involve molecular biology techniques. However, a few are available as rapid diagnostic kits for commercial
6.1 PCR tests
PCR is carried out using:
- commercial or in-house primers
- microarrays
- commercial ‘plug-in’ kits.
-
Can you name any resistance mechanisms where there are commercial PCR kits available for the detection of the resistance genes? Perhaps your laboratory uses these?
-
Examples of commercial PCR kits include:
- MecA (for S.aureus resistance identification) for rapid MRSA detection. Remember, the mecA gene is carried on a mobile genetic element present in all MRSA strains
- rapid carbapenemase tests for various genes
- you may be familiar with other commercial PCR kits that are used in your laboratory.
Benefits of commercial PCR kits include accuracy and speed. Days can be saved on turnaround times for results in the clinical setting, allowing appropriate antimicrobial treatment or infection control intervention to start earlier.
However, the negatives include:
- high cost per test
- expensive and high-maintenance equipment required
- specimens must still be cultured to look for resistance to other antimicrobials ‒ for example, if the isolate is confirmed to be MRSA, it will be β-lactam-resistant but there is no information about possible resistance to macrolides, tetracyclines et cetera.
6.2 Genome sequencing techniques
Whole genome sequencing (WGS) can also be used to look at the bacterial genome and its associated plasmids. WGS is primarily a research tool. It might be used for instance to study a newly identified mechanism of resistance or to study the different genes found in a particular setting. It could be used to understand how the resistant organisms had spread in an institution by constructing a phylogenetic tree of the bacteria in a particular species and seeing how this related to the different plasmids and resistance genes. It is also possible to sequence just the resistance genes, which is a little faster and less complex.
However, for now sequencing is still very expensive and time consuming. It also needs bioinformatics expertise to interpret the results, so even a reference laboratory is unlikely to be doing this routinely at present.
7 Genotypic versus phenotypic methods
Activity 8 Advantages and disadvantages of each approach
Complete Table 4 below then compare with the example answer.
| Comparator | Phenotypic tests | Genotypic tests |
|---|---|---|
| Speed | ||
| Equipment | ||
| Cost | ||
| Can be used for | ||
| Suitable for laboratory type | ||
| Other Advantages |
Discussion
| Comparator | Phenotypic tests | Genotypic tests |
|---|---|---|
| Speed | Culture-based tests take 24 hours Some tests, e.g. lateral flow testing, nitrocefin testing, may take only 10-15 minutes. |
Variable: PCR gives results within hours but WGS can take days to read and interpret results. |
| Equipment | Only simple requirements. | Complex – require maintenance. |
| Cost | Low – though still more than basic AST. | High – expensive equipment and maintenance. Reagents are also expensive. |
| Can be used for | Detection of mechanisms in routine clinical and veterinary practice. | Rapid PCR can detect known genes WGS can characterise new ones, but still needs to be carried out in conjunction with phenotypic testing. |
| Suitable for laboratory type | Standard clinical or veterinary diagnostic microbiology laboratory (first line). | Rapid PCR suitable for some diagnostic laboratories. Other tests done in research or reference laboratories (second line). |
| Other Advantages | Clinically useful in real time to guide antimicrobial use, infection control etc.. Does not require prior knowledge of the precise resistance genes. Can perform more tests at one time. |
Use in Epidemiology and Public health: e.g. can distinguish between mechanisms which are likely to spread and those that are not. Use of rapid PCR for hospital infection control to enable measures to be taken quickly e.g. isolating a patient with a carbapenemase-producing organism. |
8 Screening for resistance mechanisms
8.1 Theory behind screening and confirmatory testing
Many of the phenotypic and genotypic tests used to identify the resistance mechanism are time-consuming and expensive. Therefore, to reduce the total number of tests performed it is customary to screen for potential resistance mechanisms first and then confirm the mechanism on those isolates that screen positive.
The initial screening tests are done to identify organisms which may have a particular mechanism. This could involve either:
- a.routine screening of samples for asymptomatic carriage of resistance by AST, for example using samples from healthy food animals, or from asymptomatic hospital patients (for infection control purposes)
- b.testing a human or other animal clinical isolate for resistance.
Confirmatory tests then prove whether the organism has the resistance mechanism or not (EUCAST, 2017).
-
Why is the approach of screening followed by confirmation useful?
-
Screening for organisms with a resistant phenotype first followed by confirmatory tests for the resistant ones saves time and resources. You can screen large numbers quickly then focus on the resistant ones for the confirmatory tests.
Confirmation is a necessary step to avoid over-reporting of the organisms which do show resistance but do not have the mechanism of concern.

8.2 Screening for resistance mechanism in practice
Activity 9 Examples in your workplace
How might you initially screen isolates for the presence of possible resistance mechanisms? Can you give any examples from your own laboratory?
Make notes in the text box and then compare with the example answer.
Discussion
Using a specific indicator antimicrobial during routine AST of samples is a common approach to determine whether a resistance mechanism is present or not (see Section 5.1).
Examples of where this approach would be useful include:
- cefpodoxime resistance to screen for ESBL in Enterobacterales
- nalidixic acid or pefloxacin to screen for quinolone resistance in Salmonella species
- meropenem to screen for carbapenemase production in Enterobacterales (CPE).
Another approach would be to use selective media. Commercially available chromogenic media are useful especially for screening samples such as swabs or faecal samples for carriage of resistant organisms.
The choice of screening method depends on what sort of samples you are looking at, for example selective media are used for faecal samples from healthy animals, or as part of routine AST for clinical samples.
-
Chromogenic media are particularly useful if the specimen is likely to be contaminated with normal flora, for example faecal samples. Why is this?
-
Selective chromogenic media are useful if the sample has normal flora as they save time. Because only the resistant ones will be growing after overnight incubation, they can be worked on directly without having to subculture and do AST first. The test is less likely to miss the resistant ones which might otherwise be hidden among the normal flora. This method is also useful to screen large numbers of samples, and the higher cost of media is offset by savings in time.
For more information about how specific screening tests work in combination with confirmatory tests in practice, see Section 9.
9 Confirming resistance mechanisms in practice
Once screening has enabled the identification of isolates which may have the resistance mechanism you are investigating, various methods can be used to confirm whether that mechanism is present or not (EUCAST, 2017).
The following sections illustrate how confirmatory tests work in practice for ESBLs, carbapenemases and in Gram-positive organisms.
9.1 How to confirm the presence of an ESBL
9.1.1 Screening
It is important that the correct antimicrobials are used to screen for ESBLs in routine AST as only some cephalosporins are useful. If the wrong antimicrobial is used the ESBL may be missed. A combination of cefotaxime and ceftriaxone, or cefpodoxime alone will allow all types of ESBL to be detected.
-
What is the rationale behind this screening strategy?
-
Neither cefotaxime nor ceftazidime alone will pick up all the ESBLs. For example, CTX-M may look falsely susceptible to ceftazidime on disk testing but would be detected by using a cefotaxime disk. Cefpodoxime is useful because it detects all the common ones (mentioned above in section 3.1.1).
9.1.2 Synergy testing
To confirm that an ESBL is the cause of the resistance the usual approach is to test for synergy with clavulanic acid for ceftazidime, cefotaxime and cefepime, by one of several methods including:
- commercially available combination disk test (CDT) (Figure 14),
- double disk synergy test (DDST) (Figure 15)
- automated systems such as VITEK IITM or PhoenixTM .
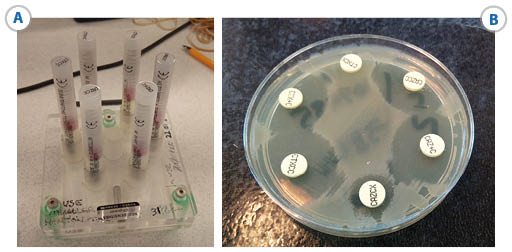
Key:
- CTX+C = cefotaxime + clavulanic acid
- CTXCX = cefotaxime + cloxacillin
- CTXCC = cefotaxime + clavulanic acid + cloxacillin
- CAZ+C = ceftazidime + clavulanic acid
- CAZCX = ceftazidime + cloxacillin
- CAZCC = ceftazidime + clavulanic acid + cloxacillin.
Activity 10 Interpreting a DDST
How would you interpret the results of the double disk synergy test in Figure 15? Does the test organism have an ESBL?
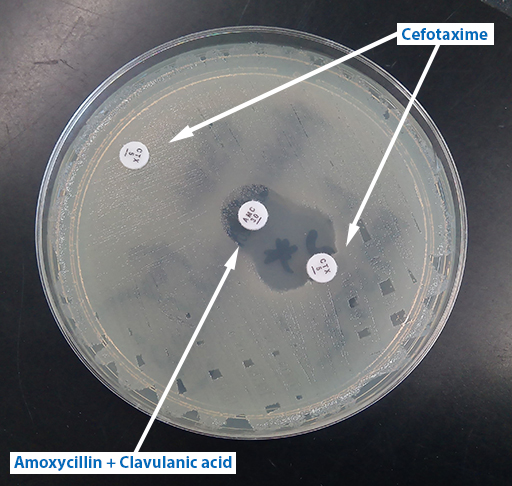
Answer
The test organism has an ESBL.
Discussion
The cefotaxime is inactivated by the ESBL (no zone) on its own but works ‒ preventing growth and leaving a clear zone – in the part of the agar where the clavulanic acid has diffused out from its disk. The clavulanic acid is protecting the cefotaxime from the ESBL. (Note the clavulanic acid disk contains co-amoxiclav (AMC) so there is also amoxicillin present. This combination has given a small zone around the disk, but not enough for this antibiotic to be useful in clinical practice.)
9.1.3 Alternatives to synergy testing
Alternatives to synergy testing include:
- colorimetric testing (biochemical)
- lateral flow test (immunological) to detect the ESBL such as NG-Test CTX-M MULTITM immunochromatographic assay
MALDI-TOF (spectrometric)- commercial PCR tests (genotypic).
9.2 Carbapenemase confirmation and identification
Meropenem is the best indicator antimicrobial to screen for carbapenemases in Gram-negatives. Resistance is implied if the MIC is raised or the inhibition zone size reduced.
It is then necessary to determine whether any resistance detected is due to a carbapenemase and not to a combination of resistance mechanisms. For example:
- ESBL or AmpC enzymes combined with decreased permeability as the result of alteration or down-regulation of porins, can also lead to carbapenem resistance
- OXA-48-like enzymes are another challenge. Organisms producing these can look susceptible to cephalosporins so it is necessary to rely on the meropenem as an indicator here. Most other carbapenemases hydrolyse all β-lactams; the OXA-48-like enzymes are unusual and easy to miss because of this unexpected sensitivity.
9.2.1 Carbapenemases in Enterobacterales
Confirmation of carbapenemase in Enterobacterales can be done by several methods in basic laboratories:
- Biochemical (colorimetric) tests, for example the CarbaNPTM test which detects the hydrolysis of carbapenem, thus confirming the presence of a carbapenemase enzyme
- Immunochromatographic tests to detect specific carbapenemases. Lateral flow tests are now becoming more readily available and are practical to use in most laboratories (Figure 16)
- For laboratories with a suitable commercial, rapid PCR platform, PCR tests for example, Cepheid, Xpert® and Carba-R, are easy to use and can identify several carbapenmases.

Other methods are possible in reference and research laboratories:
- combination disk testing – this follows the same principles as for ESBL and can potentially identify specific carbapenemases. This is more likely to be used in a reference laboratory.
- CIM (Carbapenem Inactivation Method) test and detection of carbapenem hydrolysis via MALDI-TOF are also possible, but at present do not add anything extra in the routine laboratory context.
- genotypic methods.
9.2.1 Carbapenemases in non-Enterobacterales
In non-Enterobacterales such as Acinetobacter, carbapenem resistance is due to a variety of carbapenemases with a range of porins and drug efflux pumps (Hsu et al., 2017). In Pseudomonas resistance is frequently due only to increased efflux or porin loss without a carbapenemase being present. However, carbapenemases are found increasingly in Pseudomonas and as it is hard to discriminate the mechanism in practice; the organism may need to be sent to the reference laboratory if it is important to be sure of the mechanism.
9.2.3 Carbapenemases in practice
-
From what you have learned about carbapenemases in the module, how might EDTA, a potent zinc chelating agent, be useful for identifying carbapenemases?
-
EDTA can be used to identify metallo-β-lactamases as the EDTA will remove the zinc and inactivate the enzymes. It is usually included in interpretative panels. See Section 3.1.1.
Activity 11 Identifying carbapenemases
Discussion
Note that meropenem is the best antimicrobial to screen as false positives are much less likely than with other carbapenems. For example, ertapenem could be used in Enterobacterales as it is very sensitive but might over-detect resistance leading to wasting resources in unnecessary confirmatory tests.
9.3 Confirming mechanisms in Gram-positive organisms
9.3.1 MRSA strains of S. aureus
-
Can you recall why it is important to discriminate between different types of MRSA in S. aureus, i.e. mecA derived versus hyper-production of β-lactamase?
-
Only genuine MRSA strains, i.e. those with an altered penicillin binding protein encoded by mecA or mecC genes, are of major public health and infection control importance. These are the strains likely to lead to treatment failure with β-lactam antibiotics.
Common screening methods for MRSA from swabs include colorimetric tests using chromogenic agar or mannitol salt agar. However, other tests such as the disk diffusion type tests or automated systems can be used, for example for clinical samples from invasive infections (PHE, 2020). Cefoxitin is now the recommended antimicrobial to use to confirm MRSA as it is less affected by conditions than oxacillin or methicillin and also detects mecC strains. Therefore, cefoxitin resistance confirms the MRSA phenotype.
9.3.2 Vancomycin-resistant S. aureus
Vancomycin resistance in S. aureus (VRSA) is an uncommon but increasing problem. It is worth learning how to test for it accurately.
Activity 12 Investigating possible VRSA in your workplace
You identify a S. aureus strain in your laboratory which you think may be resistant to vancomycin. For example, you may be concerned that a clinical isolate is resistant because the patient (human or animal depending on your laboratory) is not responding to treatment or, if it was collected as part of AMR surveillance, you might be doing routine testing for vancomycin resistance. What can you do to investigate this strain further?
Make notes in the text box and then compare with the example answer.
Discussion
You should be aware that disk diffusion methods are unreliable and not recommended for testing vancomycin resistance in Staphylococcus. It cannot accurately distinguish between wild type and resistant isolates.
In both situations described you need to find out the MIC of vancomycin for the isolate. This is easiest using a
Note: if you are aiming to treat a human patient or if the investigation is part of surveillance in animals with a One Health approach use EUCAST; if the aim is to treat an animal then vetEUCAST/vetCLSI guidelines (CLSI, 2020) should be used. EUCAST tables should tell you the breakpoint is 2 for Staphylococcus.
If the MIC of your Staphylococcus is vanA mechanism of resistance (e.g. modified cell wall as a result of exposure to the antimicrobial during treatment or via glycopeptide animal growth promoters used in farming). However, it is reported as susceptible for clinical and surveillance purposes.
If the MIC is >2 (resistant), then vanA is a possibility. In this case, the isolate should be sent to the reference laboratory for further testing which usually involves molecular detection of thevanA genes.
9.3.3 Vancomycin-resistant Enterococcus
When dealing with strains of Enterococcus resistant to vancomycin (VRE) the important thing is to distinguish between species that are intrinsically resistant to glycopeptide antimicrobials such as vancomycin and have no public health significance, and those which harbour acquired resistance genes.
For acquired resistance genes there are two types:
- the vanA phenotype, which are both vancomycin- and teicoplanin resistant
- the vanB phenotype which are just vancomycin-resistant.
E. faecium is the most common species to have acquired resistance and the pattern of vancomycin and teicoplanin resistance in this organism can be used to infer the presence of vanA or vanB. Gradient diffusion test strips to measure the MIC are a reliable and straightforward method to use to confirm vanA and vanB(Figure 17).
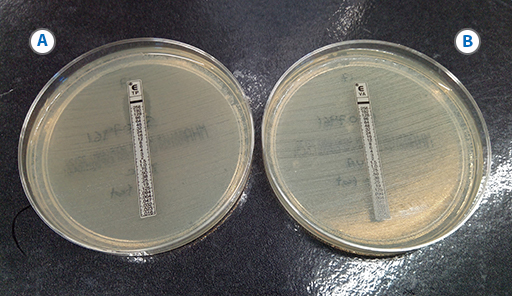
-
In Figure 17, is the test organism of phenotype vanA or vanB?
-
The organism is type vanA because it is resistant to both vancomycin and teicoplanin.
Note that disk diffusion testing is relatively unreliable for many other antimicrobials in Enterococci. It is essential to follow the guidelines very closely. Most antimicrobials do not have breakpoints for these species defined by EUCAST so it is only recommended to test the antimicrobials that do, otherwise you will end up with results that cannot be interpreted.
-
Which of the following species would need further testing for acquired resistance if they had a raised vancomycin MIC indicating resistance to this antimicrobial: E. gallinarum, E. casseliflavus, E. faecium, or E. faecalis?
-
As you learned in Section 3.2.3, E. faecium and E. faecalis would both need further testing. The others have intrinsic chromosomally encoded resistance to glycopeptides. They only rarely cause infections, for example in immunosuppressed humans. That is why these organisms are not included in the surveillance schemes.
This example illustrates how important it is to identify the Enterococcal species. However, it is not easy to tell the difference between species on biochemical tests alone. E. faecium and E. faecalis tend to be non-motile while the other species are motile and most isolates of E. casseliflavus/E. flavescens have a distinct yellow pigment, which can be observed by collecting growth from an agar plate on a swab. However, unless the laboratory has access to MALDI-TOF it may be necessary to send isolates to the reference laboratory.
9.3.4 Penicillin-resistant Streptococcus pneumoniae
Penicillin resistance of S. pneumoniae is commonly screened for with an oxacillin disk in a simple disk diffusion test. This detects non-wild type organisms if the zone size is reduced implying an increased MIC. If necessary, the MIC is then confirmed by a gradient diffusion test (Figure 18). Normally this would only be done for organisms causing an invasive infection. In general, it is seldom necessary to know the specific mechanism.
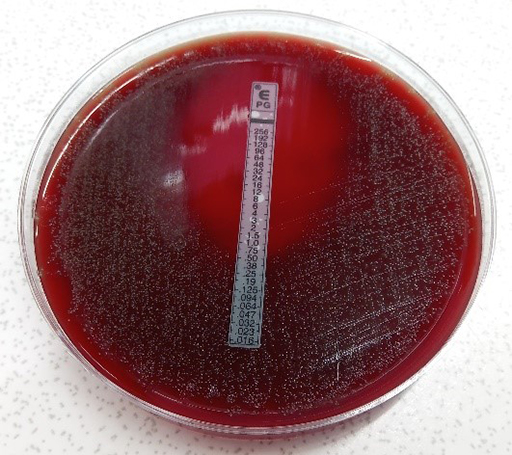
Activity 13 Interpreting an E-test to help inform a clinical decision
What is the MIC reading for the S. pneumoniae isolate in Figure 18? Refer to Table 5 and determine whether the standard dose of benzylpenicillin is suitable for treating a patient with a) pneumonia and b) meningitis.
| Penicillins | MIC breakpoints (mg/L) | |||
|---|---|---|---|---|
| S ≤ | R > | |||
| Benzyl penicillin (indications other than meningitis) | 0.06 | 2 | ||
| Benzyl penicillin (meningitis) | 0.06 | 0.06 | ||
Discussion
The pneumococcus in Figure 18 has an MIC of 0.5 mg/L. The MIC breakpoint for benzyl penicillin is
- a.according to Table 5 it is possible to use benzyl penicillin to treat pneumonia caused by this organism as the MIC is
- b.It would not be appropriate to use benzyl penicillin if this was a case of meningitis, because the blood-brain barrier reduces the amount of drug that reaches the meninges resulting in a lower level of the antibiotic in the brain than in the lungs. Therefore, a lower MIC cut-off is used for infections of the central nervous system. In this case, different antimicrobials would have to be tested to use for treatment.
How you report this isolate for surveillance will depend on the breakpoints set out in the reporting instructions.
10 How knowing the resistance mechanism contributes to AMR surveillance
10.1 The current picture
Globally, data on AMR and mechanisms are far from complete. For example, there are good data from Europe on isolates from humans and the food chain as a result of a longstanding surveillance scheme run by ECDC and EFSA, but even within these data there is variation between countries. If each centre or country only reports a few isolates this can bias the data. For countries without established regional surveillance schemes the existing data are very patchy. This makes it hard to understand the extent and distribution of resistance.
Two types of data are available:
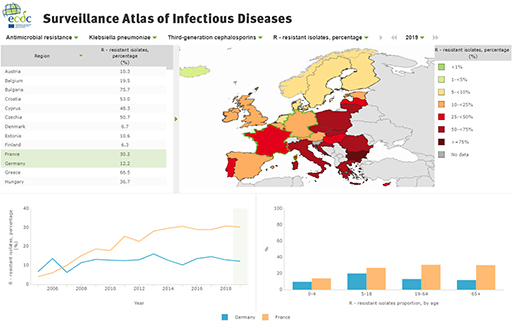
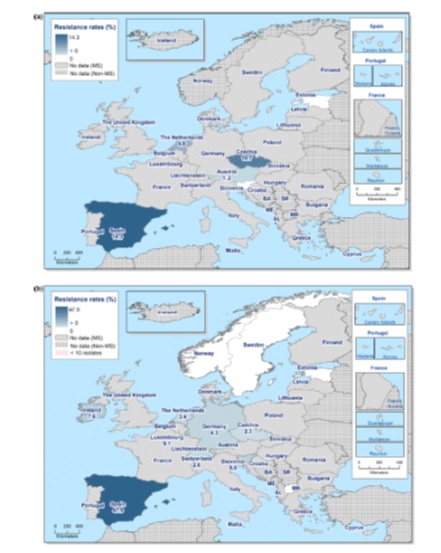
- rates of mechanisms of resistance for specific species of organisms, for example ESBLs, CPRs, VREs (Figures 19 and 20)
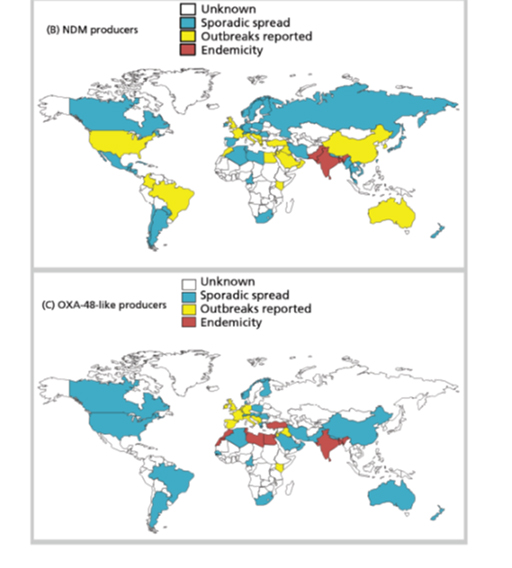
- spatial distribution of particular types of resistance mechanism, for example the NDM carbapenemases or CTX-M ESBLs (Figure 21).
These data are still however far from complete.
10.2 Importance of resistance mechanism data to AMR surveillance
Information about the resistance mechanism associated with clinical and veterinary isolates is useful at local, national and international levels.
Locally, for example within a hospital, knowing the mechanism can be helpful in order to track the spread of a particular gene and/or organism. It may also help to identify whether cases are genuinely linked. This can help the infection control team define whether it is a genuine outbreak or just sporadic cases. Additional testing would be needed to confirm whether strains are related, but it is more likely if they have the same resistance mechanism and early actions can be put in place to limit transmission. If it is an outbreak, understanding how the organism has spread makes it easier to work out the best measures to control the outbreak. For example, is it spreading from person to person or via water contamination?
On a national or international scale, data about resistance mechanisms help to identify priorities for additional surveillance and control, and for R&D. The data can enable early intervention to contain resistance mechanisms of global concern. If mechanisms can be prevented from becoming widespread, this reduces risk to patients from infections which are hard to treat and saves money in healthcare. This is even more important in low-resource settings where antimicrobial choices are limited because of financial constraints.
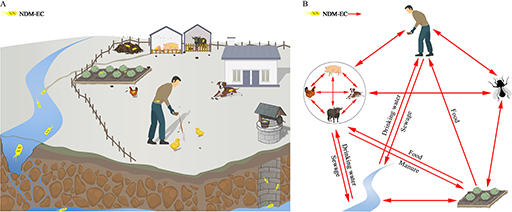
The genotypic data can also be used to help understand how transmission from food animals to the human food chain occurs (an example is shown in Figure 22, where the investigators studied the resistance genes in samples from around the farm environment). This in turn can show how this transmission can be prevented. With good quality data it is much easier to implement mechanisms to prevent transmission. This may involve complex interventions such as political, social, and financial measures which may be costly or have negative impacts on food producers in the short term so might be resisted. Data are essential to give the political drive to intervene.
Antimicrobial resistance mechanism data can also help to understand and control international transmission chains. An example could be medical tourism, where people travel to other countries either for cheaper treatment or for treatment which is not available where they live. They risk bringing back and spreading resistant strains to their home country on their return, especially if they have further hospital treatment. This is an important route of spread of AMR and is also a political issue. Again, having good data is vital in order to control spread by this route.
Finally, linking AMR resistance mechanism data to antimicrobial use is very useful to inform strategies for antimicrobial stewardship (see the Antimicrobial stewardship modules).
11 Importance of quality control
In previous modules you learned how microbiology laboratories act as ‘data collectors’ and have an important role in ensuring that test results are accurate, reliable, valid and comparable. You further learned about different
You will learn more about quality control measures in the Quality assurance and AMR surveillance module.
12 Testing for resistance mechanisms in your workplace
You have now learned about some of the more clinically important mechanisms of antimicrobial resistance and how to test for them.
Activity 14
Based on your existing knowledge and what you have learned in this module, what are the issues and challenges to bear in mind when performing tests to identify the resistance mechanism of isolates? When might you need to send an isolate to the reference laboratory for additional testing?
Make notes in the text box and then compare with the example answer.
Discussion
Some issues and challenges when identifying resistance mechanisms are listed below, but you may have thought of others:
- being aware of the most important/widespread pathogen-AMR combinations in your setting
- knowing which resistance mechanism is most likely for a given pathogen-resistance combination
- using the most appropriate screening method
- using the most appropriate confirmatory method
- understanding the principle behind common phenotypic and genotypic tests so that you are better able to recognise potential errors and anomalies
- recognising the complexity of the screening and confirmatory testing processes
- recognising that some resistance mechanisms can mask or enhance other mechanisms, e.g. AmpC and ESBL in Enterobacterales; permeability and efflux pumps in Pseudomonas
- ensuring you have training in and access to SOPs for all tests you perform
- knowing when isolates should be referred to reference laboratories for confirmatory testing. This might be if for example you cannot identify it accurately to species level, your tests of mechanism do not give you a clear answer, it is the first time you have come across this pathogen-resistance combination in your laboratory or the resistance profile is unusual for this species.
As long as your laboratory is identifying the species and mechanisms accurately, you can be confident that you are providing reliable data for surveillance. You will then be making an important contribution to fighting antimicrobial resistance both locally and around the world.
13 End-of-module quiz
Well done – you have reached the end of this module and can now do the quiz to test your learning.
This quiz is an opportunity for you to reflect on what you have learned rather than a test, and you can revisit it as many times as you like.
Open the quiz in a new tab or window by holding down ‘Ctrl’ (or ‘Cmd’ on a Mac) when you click on the link.
14 Summary
In this module you have learned about phenotypic and genotypic laboratory techniques used to test for common mechanisms of resistance in important pathogens. You have looked at resistance patterns of global concern, and you have seen how knowledge of the underlying resistance mechanism feeds into AMR surveillance. At a practical level, you have explored screening and confirmatory tests used for pathogen-resistance combinations relevant to your workplace.
You should now be able to:
- give examples of the resistance patterns/resistant organisms causing global concern
- give examples of the resistance patterns/resistant organisms encountered in your work
- understand the difference between screening and confirmatory testing
- describe some of the phenotypic methods commonly used for screening and confirmation of resistance mechanism
- understand how genotypic methods can be used, and the advantages/disadvantages of genotypic versus phenotypic methods
- understand how more data from detailed testing for resistance mechanisms/genes contributes to AMR surveillance
- apply your knowledge of these laboratory tests to interpret data relevant to your work
- know the importance of procedures designed to ensure the quality of these laboratory tests in your workplace.
Activity 15 Reflecting on your progress
Do you remember at the beginning of this module you were asked to take a moment to think about these learning outcomes and how confident you felt about your knowledge and skills in these areas? Now that you have almost completed this module take some time to reflect on your progress and use the interactive tool to rate your confidence in these areas using the following 1-5 scale.
- 5 Very confident
- 4 Confident
- 3 Neither confident nor not confident
- 2 Not very confident
- 1 Not at all confident
Try to use the full range of ratings shown above to rate yourself.
References
Acknowledgements
This free course was collaboratively written by Sarah Palmer and Liz Sheridan, and was reviewed by Priya Khanna, Skye Badger, Claire Gordon, Natalie Moyen and Hilary MacQueen.
Except for third party materials and otherwise stated (see terms and conditions), this content is made available under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 Licence.
The material acknowledged below is Proprietary and used under licence (not subject to Creative Commons Licence). Grateful acknowledgement is made to the following sources for permission to reproduce material in this free course:
Images
Module image: © funtap/123RF
Figure 1: EYE OF SCIENCE/SCIENCE PHOTO LIBRARY
Figure 2: The Open University
Figure 3: Liz Sheridan
Figures 4 and 5: Liz Sheridan
Figure 6: © Anant Kasetsinsombut/123 Royalty Free
Figure 7: Liz Sheridan
Figure 8: EYE OF SCIENCE/SCIENCE PHOTO LIBRARY
Figures 9, 10, 11 and 12: Liz Sheridan
Figures 13, 14, 15, 16, 17 and 18: Liz Sheridan
Figure 19: © ECDC. Dataset provided by ECDC based on data provided by WHO and Ministries of Health from the affected countries
Figure 20: EFSA and ECDC (European Food Safety Authority and European Centre for Disease Prevention and Control), 2021. The European Union Summary Report on Antimicrobial Resistance in zoonotic and indicator bacteria from humans, animals and food in 2018/2019. EFSA Journal 2021;19(4):6490, 179 pp. https://doi.org/10.2903/j.efsa.2021.6490. This file is licensed under the Creative Commons Attribution-NoDerivatives Licence http://creativecommons.org/licenses/by-nd/4.0/
Figure 21: Bonomo, R.A., Burd, E.M., Conly, J., Limbago, B.M., Poirel, L., Segre, J.A. and Westblade, L.F. (2018) 'Carbapenemase-Producing Organisms: A Global Scourge', Clinical Infectious Diseases, Vol. 66, Issue 8, 15 April 2018, Pages 1290–1297. Oxford University Press.
Figure 22: Li, J. et al. (2019) Inter-host Transmission of Carbapenemase-Producing Escherichia coli among Humans and Backyard Animals, Environmental Health Perspectives, Vol. 127, No. 10, https://doi.org/10.1289/EHP5251
Videos
Video 1: Liz Sheridan
Tables
Table 1: With kind permission of Prof Peter Hawkey, personal communication, University of Birmingham
Table 5: European Committee on Antimicrobial Susceptibility Testing (EUCAST) (2021) ‘Clinical breakpoints – bacteria’, EUCAST.
Every effort has been made to contact copyright owners. If any have been inadvertently overlooked, the publishers will be pleased to make the necessary arrangements at the first opportunity.