Use 'Print preview' to check the number of pages and printer settings.
Print functionality varies between browsers.
Printable page generated Tuesday, 21 May 2024, 2:46 PM
Diagnostic stewardship in clinical practice
Introduction
Diagnostic stewardship is a systematic approach to the effective use of the microbiology laboratory in clinical practice in order to deliver safer, more effective and efficient patient care by appropriate and timely generation of clinically relevant microbiological data.
The primary aim is to benefit the patient and ensure that they receive the correct treatment. Proper use of diagnostics also helps to understand the prevalence of pathogens in general, and AMR in particular. An appropriate approach to testing generates accurate and representative AMR surveillance data to inform local and national treatment guidelines and AMR control strategies. Furthermore, data resulting from its systematic use can contribute to pathogen-focused AMR surveillance such as the the World Health Organization’s (WHO’s) Global Antimicrobial Resistance Surveillance System (GLASS).
This module will introduce diagnostic stewardship with a focus on the essential structures and practices in both clinical and laboratory settings, and the role of administrators in enabling effective collaboration and data collection.
After completing this module, you will be able to:
- describe the roles in a diagnostic stewardship programme
- understand the principles of taking appropriate clinical samples
- appreciate the range of laboratory techniques available for bacterial isolation, pathogen identification and AST
- understand how microbiology can be appropriately reported to clinicians
- understand factors affecting bacteriology laboratory turnaround times and reporting
- promote good working relationships between laboratories and clinicians for effective diagnostic stewardship
- understand diagnostic stewardship in surveillance at national and international levels
- understand how to introduce a diagnostic stewardship programme.
Activity 1: Assessing your skills and knowledge
Before you begin this module, you should take a moment to think about the learning outcomes and how confident you feel about your knowledge and skills in these areas. Do not worry if you do not feel very confident in some skills – they may be areas that you are hoping to develop by studying these modules.
Now use the interactive tool to rate your confidence in these areas using the following scale:
- 5 Very confident
- 4 Confident
- 3 Neither confident nor not confident
- 2 Not very confident
- 1 Not at all confident
This is for you to reflect on your own knowledge and skills you already have.
1 Diagnostic stewardship and managing AMR
A
1.1 Where does diagnostic stewardship fit in?
In the context of
Studies suggest that AMS programmes are more effective when coupled with
1.2. Saving money, improving patient outcomes
As more tests become available, including a multitude of rapid diagnostic tests (RDTs), choosing the appropriate test for a particular patient in a particular setting requires the combined knowledge and expertise of clinical microbiologists, technicians, pharmacists, data analysts and clinicians.
Without these experts’ active collaboration, a clinician could order unnecessary tests or fail to request the most appropriate one. A 2013 study estimated that 20% of tests were overused and approximately 45% of tests were underused (Zhi et al., 2013). Although the tests are less than 5% of a hospital’s total costs, inappropriate use of tests may lead to unnecessary interventions or treatments that may be harmful to the patient, and/or cause a failure to provide appropriate treatment.
Both of these factors are detrimental to the patient but may also lead to longer hospital stays, the cost of which can dwarf the cost of laboratory testing. Successful diagnostic stewardship has been shown to effectively reduce a variety of unnecessary tests, from excessive or redundant daily inpatient laboratory tests to diagnostic imaging (Dik et al., 2017).
As more tests (including RDTs) become available, it becomes more difficult for the clinician to choose the appropriate test without advice from a clinical microbiologist, and more difficult for the laboratory to know which tests to add to their list without specific input from the clinicians and the pharmacy. Information on local prescribing practices and prevalence of specific infections and resistance patterns is also required.
As well as determining which tests are appropriate, diagnostic stewardship also encompasses correctly collecting and managing samples to allow a reliable result that is reported in a timely and effective manner to the clinician, and is interpreted correctly.
Activity 2: Your experience of diagnostic stewardship
Think about the situation in your hospital or healthcare setting.
- Do you think microbiological tests are overused or underused at your facility?
- If you are not sure about this, then if you are a clinician, consider whether you have a full appreciation of all the available tests and their correct use.
- If you are a clinical microbiologist or laboratory scientist, do you have the opportunity to provide your input to the clinicians ordering the tests? Do you have a good knowledge of local patterns of antimicrobial use (AMU), which may include over-the-counter sales outside the hospital, and AMR?
- If you are at the administration or analysis level, do you think the tests available at your site(s) give you all the information you need?
- What simple changes could you make to your practice to ensure that appropriate tests are ordered and the results interpreted correctly?
Discussion
You may feel that tests are being used appropriately, but that other tests are currently unavailable at your hospital laboratory. Some of these tests could be beneficial, and this opportunity is something you could discuss with representatives from clinical, laboratory and pharmacy sectors, together with management and epidemiologists.
You could have suggested that access to the laboratory is important: transportation of specimens may be an issue in a centralised laboratory system.
If you are a clinician, you may think that you are ordering tests because you want to cover all possibilities or because you have always ordered these tests, but that you would benefit from more guidance at this stage from the laboratory.
Perhaps you suggested that a committee could be formed, comprising individuals with the relevant expertise – such as infectious disease clinicians, clinical microbiologists, administrators, pharmacists or epidemiologists– to plan and implement improvements. This is a key requirement in implementing diagnostic stewardship!
1.3 Finding the ‘sweet spot’
Although reducing the number of unnecessary tests has many potential benefits for the patient and hospital, underuse of tests could result in serious infections going undiagnosed and untreated. One of the major objectives for diagnostic stewardship for AMR is to identify the ‘sweet spot’: reducing over-diagnosis and false positive results while benefitting from appropriately indicated testing and true positive results. Where this spot lies is infection- and population-specific: that is, related to disease prevalence.
For example, the use of urinalysis dipstick tests and urine culture in elderly female patients without signs and symptoms of a urinary tract infection (UTI) is debatable. These tests are often performed in patients with non-specific presentations (such as a fall or dizziness), and the symptoms are attributed to a UTI, with antimicrobials being prescribed. However, it is known that in this age group, patients often have low-level bacteriuria (bacteria in the urine) that, while not sufficient to cause a UTI, will result in a positive test. The patient will then receive unnecessary antimicrobials and the actual cause of their symptoms might not be properly investigated.
Another example comes with testing for Clostridioides difficile (C. difficile). Although C. difficile is an important infection (associated with AMU), a small proportion of patients carry C. difficile in their gastrointestinal tract without having active infection. Testing may therefore identify patients with colonisation or with resolving infections. There is a fine balance between colonisation and invasion by microorganisms; clinical assessment is required to determine whether the
A patient benefits from clinicians and microbiologists liaising with each other. If the patient has had a recent infection with C. difficile, there is no benefit to testing: we would still expect the test to be positive, and so it would be a waste of resources. Similarly, a patient with a positive test needs to be assessed to see if they have signs and symptoms of infection rather than colonisation, to prevent unnecessary treatment.
All of this must be balanced with the knowledge that C. difficile infection can be very severe, and is readily transmitted to other patients. Restricting testing too much might result in unrecognised and untreated C. difficile infection, which would result in harm to individual patients and a greater risk of cross-infection. This example demonstrates that providing a test also requires an appreciation of its appropriate use: this is the rationale for improving diagnostic stewardship (Baron et al., 2013).
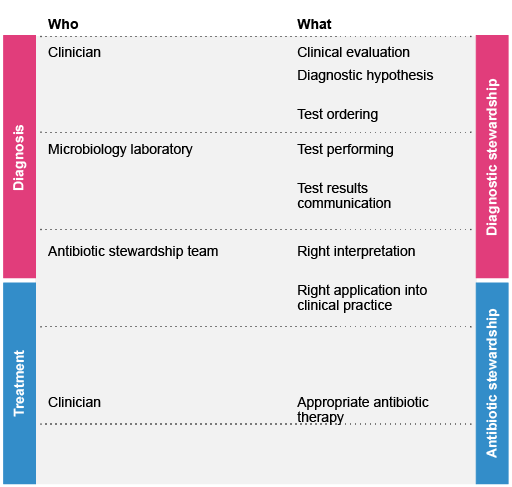
2 The diagnostic workflow in the clinical microbiology laboratory
Clinicians make diagnoses based on the patient’s history and risk factors, physical findings on examination and the results of diagnostic testing (such as imaging, laboratory tests, etc.).
Laboratory diagnostic testing can be broken down into three stages:
- Pre-analytical: Test-related decision-making and specimen collection.
- Analytical: The test itself and any associated laboratory practices, including test methods, microscopy, culture and identification, antimicrobial susceptibility testing (AST), and protocol validation.
- Post-analytical: Reporting and use of laboratory data, such as selective reporting of antimicrobial susceptibility data to encourage the use of narrower spectrum agents.
From the clinical perspective, diagnostic stewardship focuses on stages 1 (pre-analytical) and 3 (post-analytical); most clinicians are not involved in the actual test performed in laboratory (stage 2). The overall value of the test itself relies on good quality at all stages, not just at the analytical stage.
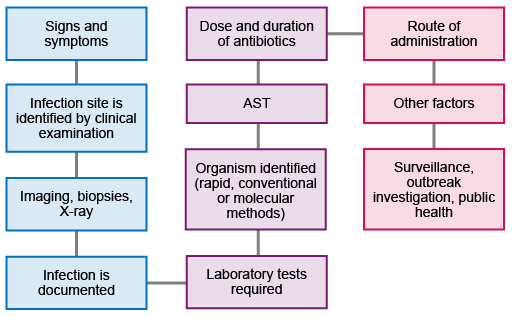
Figure 2 shows the workflow from initial presentation of the patient through to the collection and use of the data to inform further action and strategy. Not all patients will require all the steps: for example, not everyone will require imaging or biopsies, and not all infections will need public health interventions.
Key processes of laboratory diagnostic testing in the pre-analytical, analytical and post-analytical steps are shown in Table 1.
| Stage | Process |
|---|---|
| Pre-analytical |
|
| Analytical |
|
| Post-analytical |
|
Sections 3–5 will consider the three stages in more detail. The personnel involved in each stage will depend on the institution, but will usually include physicians, laboratory staff, infection prevention and control teams, and pharmacists. Where available, clinical microbiologists (medical doctors who have specialised in microbiology) should oversee and coordinate the pathway.
3 The pre-analytical stage
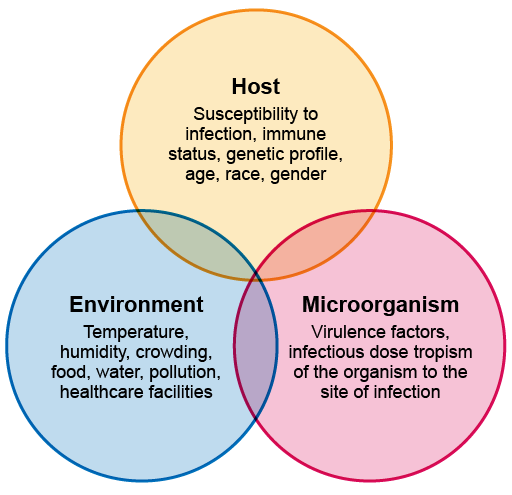
The clinical presentation of an infectious disease reflects the interaction between the host, the microorganism and the environment. When they are assessing a patient, the clinician – who could be a doctor, clinical officer, nurse, physician associate, community healthcare worker, etc. – needs to consider:
- the patient’s history and reported symptoms
- any physical findings
- any risk factors that might pre-dispose them to particular infections, such as whether they are immunosuppressed, what diseases are prevalent locally or any occupational exposures.
A clinician must also consider any local outbreaks of diseases such as cholera, or disease outbreaks in healthcare facilities related to antimicrobial-resistant organisms. For AMR, a history of recent healthcare exposure and whether or not the patient has recently taken antimicrobials is also important, because these are risk factors for resistant infections.
Signs and symptoms also vary according to the site and severity of infection; they may be localised or systemic, with fever, chills and hypotension. For example, patients with an uncomplicated UTI may have localised symptoms such as pain on passing urine; but if the infection is more severe and involves their kidneys, they may have back or loin pain, fever and chills, weakness and tenderness, and are likely to be unwell. Patients with bloodstream infections are likely to be very unwell, with systemic fever and low blood pressure.
The origin of the infection also needs to be considered.
Figure 3 gives an overview of these interactions.
Although this modules focuses on bacterial infections, you should remember that patients presenting with fever may have other types of infection, and the history and examination should consider all the possible infection types. Again, the patient’s exposure history and information about local
3.1 What affects appropriate sample selection?
Diagnosis requires a composite of information, including history, physical examination, radiographic findings and laboratory data. Specimens are selected based on signs and symptoms.
It is important that samples should be representative of the site of infection, and should be collected before antimicrobial agents are administered (otherwise the growth of organisms in the laboratory will be suppressed). You should also consider that the patient may have taken antimicrobials acquired outside the healthcare system before presenting.
As noted above, other types of infection also need to be considered. In some cases, sampling follows a protocol or algorithm, where patients with particular symptoms are initially tested for one type of infection (such as malaria, or Covid-19); if these are negative, or if the patients do not respond to initial treatment, you can then make further investigations for other infections or non-infectious conditions. Clinical microbiologists or infectious disease specialists can provide advice on prioritising tests or developing testing algorithms.
3.2 The principles of sample collection
Collecting the sample requires great attention to detail. Education and training should be available to clinicians to ensure that they collect samples that will provide useful and accurate laboratory results.
It is critical to avoid contamination with normal microbial flora from the patient, and from the person taking the sample: otherwise, treatment might be misdirected. Avoiding contamination is particularly important for cultures of blood, bone and other tissues or fluids in which infection is often caused by indigenous flora, and for specimens collected from sites of putative infection that are contiguous to or adjacent to cutaneous or mucosal surfaces. It is therefore necessary to use strict
Activity 3: Collecting blood culture samples
Watch the video ‘Prof Koch’s guide to perfect blood cultures’, which discusses the collection of samples for blood culture. Then complete the two tasks below.
- Make a note of the details of appropriate blood culture collection procedure mentioned in the video.
- How do you think your hospital could ensure blood culture samples are taken correctly?
Discussion
- You may have noted the following requirements:
- When testing for bacteraemia, try to take a sample as the temperature begins to spike
- Take the blood culture sample before administering antimicrobials
- Avoid the femoral vein
- Avoid existing cannulae and lines
- Bottles must be in date and in good condition
- Disinfect site with 2% chlorhexidine with 70% isopropyl alcohol
- Allow the site to dry thoroughly after disinfection
- Don’t touch the skin again after disinfection
- Swab the rubber tops of the bottles with ethanol, and allow to dry
- Inoculate aerobic bottles before anaerobic bottles
- Collect as much blood as possible
- Discard the needle directly into the sharps box to avoid needle stick
- The bottle must be correctly labelled
- Keep bottles at room temperature
- Note that there are alternatives for disinfection, such as 10% povidone-iodine (see Table 2).
- You may have noted the following:
- The laboratory could provide a checklist for samples and refuse to test samples submitted without full details or otherwise incorrectly managed.
- Posters could be put up in wards showing the agreed correct procedure, using diagrams.
- Regular refresher training could be provided to healthcare staff taking blood culture samples.
- Note that there are different options for the procedure, so details of these can be agreed by discussion between the clinical and laboratory sectors.
3.3 Handling and transporting specimens correctly
Generally, samples should reach the laboratory with no delay, and ideally within two hours. Prolonged transportation may result in the death of fastidious bacteria and overgrowth/colonising of other bacteria.
The hospital administration may need to organise appropriate courier services to ensure that samples arrive in the laboratory within the required time if the laboratory is not close to the hospital. If it is necessary to store a sample, appropriate storage facilities such as fridges need to be made available and accessible. The clinical and laboratory administration may need to discuss who should take responsibility for maintenance and security of storage facilities and transport.
The laboratory needs to have reception facilities and procedures (including verification that a sample is appropriate), and should immediately inform the clinician if the sample is not adequate, explaining the reasons so that the sample can be collected again. Inappropriate samples should be rejected, since they will give no useful information and could provide misleading information. Local
3.4 Guidance for collecting and transporting specific samples
This section includes more detailed information on common specimens, including blood, cerebrospinal fluid (CSF), respiratory samples, faeces and urine (Tables 2–6).
| Collection | Only a proportion of blood cultures yield significant microorganisms. The proportion will depend on who is sampled and local infection prevalence, but usually, fewer than 10% of samples will grow significant pathogens. Even with good collection technique, 1–3% of blood cultures are contaminated. Blood culture contamination rates are minimised by strict adherence to aseptic collection technique and, whenever possible, collecting peripheral blood via venepuncture with proper skin antiseptic preparation, for example with 70% alcohol, 0.5% chlorhexidine gluconates or 10% povidone-iodine. |
| Volume | The total volume of blood for blood culture request is the most important aspect, because bloodstream infections have a low concentration of organisms in the blood, estimated as 99% of pathogens, but in practice it is very difficult to obtain this volume. For optimal sampling in adults, it is recommended that 20–30 ml of blood is taken per set (a set means an aerobic bottle plus an anaerobic bottle), and that two sets (four bottles in total) are taken. While this is ideal, local decisions may need to be made based on sampling costs. For suspected infective endocarditis, three sets of blood cultures are recommended, taken from different sites one to two hours apart. This is because endocarditis may be caused by normal skin flora (especially prosthetic valve endocarditis) and because patients will need to be on a long course of antibiotics. It is therefore vital to make sure that the diagnosis and causative organism are correctly identified: the results of three or four positive blood cultures with the same organism establishes the presence of continuous bacteraemia and helps the physician determine the clinical relevance of the isolates. In children, the optimal volume of blood is less well-prescribed; and especially for infants, the need for volume to improve test sensitivity needs to be balanced with the risk of over-bleeding the infant. The recommended volume is therefore based on the weight of the child: for infants less than 1–2 kg, 2 ml should be submitted; for a child of 2–12 kg, 4 ml; and more than 12 kg, 10 ml. A single aerobic bottle should be used unless anaerobic infection is suspected. Alternatively, a single special paediatric culture bottle can be used. If a vascular line infection is suspected, cultures should be taken from both the line and peripherally, to determine whether the line is contaminated or infected, and whether the infection has become systemic. |
| Transport | Specimens should reach the laboratory with no delay. Although commercial blood culture bottles have been carefully formulated to optimise bacterial growth, prolonged transportation may result in death of fastidious bacteria and overgrowth of other bacteria. If there is going to be a long delay in sending specimens to the laboratory, the bottles should be kept in an incubator at 35–37°C in an onsite laboratory or in a safe place on the ward. |
| Collection | Cerebrospinal fluid (CSF) must be collected using strict aseptic technique, both to minimise specimen contamination and to prevent bacteria being introduced into the central nervous system. Either povidone-iodine or chlorhexidine can be used for disinfection. The risk of contamination is higher when CSF is collected from catheters or shunts. Such contamination is problematic, because organisms (such as coagulase-negative Staphylococci) are likely to cause many CSF catheter and/or shunt infections: it may therefore be necessary to take several samples to help to distinguish contamination from true infection. |
| Volume |
The volume of CSF depends on the pathogens sought. For routine bacterial cultures, 2–3 ml is adequate for adults. Note that for fungal cultures, microbial yield is more proportional than bacterial yield to the volume of cerebrospinal fluid cultured. The laboratory should consider using sequential testing to reduce the number of unnecessary CSF tests, for example by use of a testing algorithm that specifies a chronological order that tests should be performed in. For example, the first step might be recording opening pressure to assess if inflammation is present, followed by assessment of colour, or Gram staining for bacterial meningitis. (The ddxof website provides an example of an algorithm for analysing CSF.) When testing CSF for tuberculosis (TB) culture, samples can be sent for acid-fast smear with the important caveat that a single sample (approximately 20–40%) has low sensitivity. Several large volume (10–15 ml) lumbar punctures are often needed for a microbiological diagnosis; sensitivity increases to >85% when four spinal taps are performed. While culture can take several weeks and has low sensitivity (~40–80%), it should be performed to determine drug susceptibility (Marx and Chan, 2011). |
| Transport | CSF specimens should be transported immediately to the laboratory. For suspected bacterial meningitis, the laboratory should report the results of initial tests (including microscopy with cell counts and Gram stain, and latex agglutination for bacterial organism such as N. meningitidis) within 30 minutes of receipt of the specimen so that appropriate treatment can be given. From collection through processing, CSF specimens (except aliquots collected for viral cultures, performed only in specialised centres) should not be refrigerated until initial processing is completed. |
| Collection | For diagnosing lower respiratory tract infections (LRTIs), expectorated sputum is the most commonly received sample, as it can be obtained easily and non-invasively. However, patients with pneumonia often have difficulty in producing a good-quality sputum sample due to pain and breathlessness, and specimens are frequently contaminated by normal resident bacteria of the oropharynx, preventing the determination of the true pathogen and leading to wrong results. More invasive samples (such as biopsies, brushings and lavage specimens) have better sensitivity and specificity, but require bronchoscopy, which is rarely required or performed for uncomplicated pneumonia. The value of sputum microscopy and culture in the diagnosis, management and outcome of LRTIs is therefore a matter of controversy, and many laboratories no longer process sputum samples routinely. |
| Volume | The quality of an expectorated sputum sample can be assessed by Gram staining to look for epithelial cells (indicative of a poor quality sample), white blood cells and organisms (indicative of infection). The average number of the different cell types in 20–30 low power fields is calculated and the total score is based on Bartlett criteria. A final score of 0 or less indicates lack of active inflammation or contamination (non-acceptable sample), and a score of 1 and above is considered an acceptable sample. Only samples with a score of 1 or above should be cultured. A sputum Gram stain examined according to the correct guidelines is considered useful in the initial evaluation of patients with pneumonia (Del Rio-Pertuz et al., 2019). However, it is labour-intensive, and local guidelines should be developed to optimise the cost-effectiveness of processing sputum samples. An additional factor to consider is the local prevalence of TB, as there may be a risk of transmission to laboratory staff from sputum samples and appropriate precautions should be taken. |
| Transport | Most respiratory tract specimens are likely to contain at least a few contaminating microorganisms, so specimens should be transported rapidly to the laboratory to minimise contaminant growth. If transportation or processing is delayed for more than two hours, specimens should be rejected. |
| Collection | The laboratory diagnosis of enteric infections is challenging, due to the diversity of the normal flora. Local policy should decide which organisms should be tested for. GLASS (which will be introduced later in this module) looks for Salmonella spp. and Shigella spp. due to the emergence of multidrug-resistant strains, but many laboratories will also look for Campylobacter spp. Stool specimens collected from patients who develop diarrhoea in the hospital should be tested for the presence of C. difficile toxin. |
| Volume | There is little value in routinely testing multiple stool specimens as part of an evaluation of acute diarrhoea, as most pathogens are detected in the first specimen. Repeat specimens should be sent if symptoms persist. |
| Transport | Samples should be transported to the laboratory within two hours. Shigella spp. are very sensitive, so prompt delivery to the laboratory is critical if they are to be cultured successfully (WHO, 2016). |
| Collection | Because urine is easily contaminated with commensal flora, careful collection of specimens is vital to minimise this. Midstream urine is ideal; first morning urine is best because it may contain more pathogenic organisms. Patients need to be instructed on how to clean themselves and take the midstream urine. Samples from catheter tips and bags are not appropriate (Miller et al., 2018) because they are frequently colonised and/or contaminated. |
| Volume | The volume of urine specimens is not as important as collecting urine in sterile specimen containers, and obtaining a ‘clean catch’ or midstream urine. |
| Transport | Urine samples should be collected and reach the laboratory as soon as possible, and culture should commence immediately to avoid overgrowth of other non-pathogenic organisms. Specimens that won’t reach the laboratory within two hours of collection can be refrigerated for up to 24 hours. Preservatives such as boric acid, which inhibits bacterial growth and prevents overgrowth with contaminants, may be used for storage and transportation of urine. Non-buffered boric acid is preferred so as to reduce the effect of the preservative on the actual pathogens (Weinstein, 1983). |
3.5 General requirements for specimen processing
The following general considerations for specimen processing are relevant for all sample types processed for bacterial culture, identification and AST:
- Poor-quality specimens, or specimens that are poorly transported – for example, in a leaking container – should be rejected. A local SOP should be developed for this.
- Specimens must be labelled appropriately, with three identifying data points (such as name, date of birth and hospital number) and should match the request form. The time and date of collection should also be indicated.
- The request form should include relevant clinical information, including the suspected diagnosis, which may help the laboratory decide on sample processing, and whether any additional precautions should be taken.
- Normal flora should not be reported.
- Strict protocols are required for submission of specimens from non-sterile sites, such as pus from open wounds.
- Laboratory procedures must require appropriate volumes of specimens to maximise recovery of the pathogens. Clinicians should be aware of the optimal volumes for different samples.
- Laboratories must follow strict laboratory SOPs.
- Specimens must be collected prior to antimicrobial administration, unless taking the sample would delay administration of emergency antimicrobials. For example, if the initial attempt at lumbar puncture for suspected meningitis was unsuccessful and the patient was very unwell, it would not be appropriate to delay antimicrobials any further.
- AST should be reported for all clinically relevant isolates.
- Microbiology reports must be clinically relevant.
- The laboratory and medical staff should develop SOPs and technical policies relating to the submission of specimens.
Laboratory personnel receive a urine specimen collected 24 hours prior in a container that has leaked, leaving only a drop of urine. On the label, the patient’s name is unclear, but it is assumed that they have received initial empirical treatment. What recommendations should laboratory staff follow regarding management of this specimen?
-
- A specimen of poor quality or that is poorly transported, such as in a leaking container, should be rejected.
- The specimen must be labelled appropriately and completely; if not, it must be rejected.
- Strict instructions for submission of urine specimens must be in place and followed.
- The specimen must be collected prior to antimicrobial administration, if possible.
- Urine specimens must be refrigerated if transport to the laboratory is not possible within two hours.
- The microbiology laboratory should send a report about the rejected specimen to the clinician and ask for a repeat sample if clinically indicated. For critical specimens (such as CSF samples), the clinician should be contacted immediately.
The laboratory should provide an easily accessible list of tests available to all users, including guidelines, turnaround times and other information on their relevance. Ideally this should be available in an electronic format so that it can be easily updated.
4 The analytical stage
Diagnostic techniques range from conventional culture-based identification and AST to molecular diagnostic methods. (These techniques are discussed in detail in the modules Isolating and identifying bacteria and Antimicrobial susceptibility testing, so are not explained here.) Each technique has advantages and disadvantages, and a basic understanding is useful to help clinicians optimise their use of the laboratory.
Culture-based methods are part of routine pathogen detection. They require viable organisms to culture, an incubation period for growth and more than 24 hours for results: this means it can take two to three days between taking a sample and getting the final result – sometimes longer for difficult-to-grow bacteria.
More rapid techniques are available: such as quantitative PCR (qPCR), which can quantify the amount of pathogen genetic material and determine the presence of specific genes and alleles. However, these should be used with care, because genes present are not necessarily expressed, and do not necessarily correlate with phenotype. The rapid test may be useful for initial treatment, but results should be correlated with phenotypic and biochemical tests (Kralik and Ricchi, 2017). qPCR can also be used for typing strains and isolates, and for indicating potential toxin production.
Validated international guidelines are available for standard culture techniques and AST such as disk diffusion and broth microdilution. If these are followed carefully, laboratories can have high confidence in their results, and data can be readily compared between facilities and geographic locations. For AST, the international standards used are CLSI and EUCAST (see the module Antimicrobial susceptibility testing).
4.1 Rapid diagnostic tests
Savoldi et al. (2020) reviewed the use of rapid diagnostic tests (RDTs) in conjunction with close collaboration between clinicians and clinical microbiologists. They found that integrating RDTs into the diagnostic workflow requires knowledge of:
- the local epidemiology of phenotypic resistance
- the local epidemiology of molecular resistance mechanisms, if available
- the general or local empiric treatment guidelines.
RDTs improved the time to result and – despite the initial investment in equipment – saved money in the longer term, particularly if combined with AMS. Timely communication between clinicians and the laboratory enabled the appropriate use of tests and real-time notification of test results. In various studies reviewed by Savoldi et al. (2020), this resulted in the earlier transition from empirical to specific treatment, better patient outcomes and a reduced length of stay in hospital – which consistently resulted in an overall cost saving per patient.
Nucleic acid-based rapid tests are commercially available and are easy to carry out. They can be a rapid way to identify pathogens and antibiotic-resistant genes (ARGs). Often they are multiplexed to look for several pathogens and ARGs at the same time, and may return results in as little as one hour. Examples include the Verigene Gram-positive blood culture nucleic acid test, which can be used directly on blood culture bottles, in conjunction with other diagnostic tests, and the FilmArray™ blood culture identification (BCID) panel, which tests for 24 pathogens and three ARGs associated with bloodstream infections.
For RDTs, as for other tests in microbiology, interpretation by the clinical microbiologist is essential, and the test result must be considered in the context of the clinical information and patient history. Note that some RDTs available to support the diagnosis of infectious diseases do not necessarily identify the pathogen, but may indicate an immune response to infection such as the test for CRP (C-reactive protein). The size of the increase in CRP helps to distinguish bacterial infection from viral infection.
Unlike RDTs available for other infections such as malaria, the RDTs currently available specifically for bacteria still require the sample to be processed in the laboratory, and usually, bacteria have to be isolated and grown on an agar plate first – so it may take 18–24 hours before the result is available. RDTs for bacteria are therefore not yet available for near patient or bedside testing.
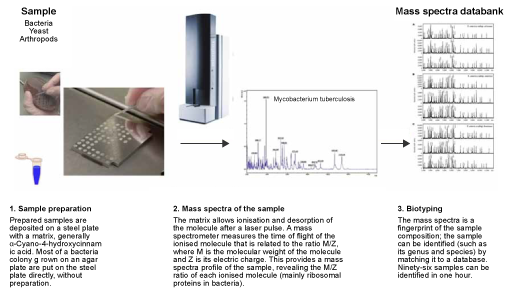
4.2 MALDI-TOF mass spectrometry
MALDI-TOF mass spectrometry (MS) is a specialised type of laboratory RDT. Most laboratory RDTs can only look for one particular type of bacteria, and standard identification requires different processes, reagents and equipment for each different type of bacteria. MALDI-TOF MS uses the same process (identification of bacterial protein profiles according to their molecular weight and charge) for all types of bacteria, using the same reagents.
Once bacteria are visible on an agar plate (usually after 8–18 hours of incubation), their species can be identified using MALDI-TOF MS in a matter of minutes, whereas standard testing requires a laboratory technician to set up a series of tests (which is itself time-consuming) and incubate for a further 24–48 hours.
The cost of reagents is very small, but a MALDI-TOF mass spectrometer is a significant investment: the prospective cost of maintenance and a database that needs regular updates should be considered. Usually, the investment needed to purchase and maintain a MALDI-TOF is only worthwhile in laboratories processing tens of thousands of samples per year, or in reference laboratories.
4.3 Choosing laboratory tests
As noted above, the use of RDTs requires close collaboration between clinicians and microbiologists, and decisions about introducing new tests may be made by a diagnostic stewardship committee. This committee will be aware of the long-term cost savings and patient benefits of using RDTs. Larger facilities may have a clinical microbiologist, who will advise on new tests and oversee their evaluation and introduction.
Diagnostic tests must be appropriate for the individual patient and should target all pathogens that cause acute infections.
4.4 Laboratory and clinician communication
Helping individual physicians to select and interpret diagnostic tests on the appropriate clinical specimens is the major goal of diagnostic stewardship, and good communication between the laboratory and the clinicians is important in achieving this. Similarly, clinicians can help laboratories to process samples optimally by providing relevant patient clinical data and clinician contact details on the request form. The laboratory should inform the clinician when they can expect to receive the result, and should notify the clinician with the result, explaining its interpretation (if needed) and ideally being available by phone to answer any questions (Dik et al., 2017).
5 The post-analytical stage
Now it’s time to look at the final stage of laboratory diagnostic testing.
Before exploring the post-analytical stage in more detail, can you recall the critical roles of the microbiology laboratory?
You may have suggested that the critical roles of the microbiology laboratory are to:
- help individual physicians to select the appropriate test for their patient
- provide right, rapid, relevant and reliable results to diagnose infectious diseases
- explain the report in the context of the medical background, and the microorganisms involved in infection – it is necessary to interpret the data in context of the clinical condition of the patient
- report results to clinicians, to influence their infection management and antibiotic prescribing behaviour (e.g. selective reporting).
5.1 Reporting results immediately and appropriately
Rapid notification of even preliminary blood culture results, such as growth and Gram stain results, can impact rational antibiotic prescriptions, length of hospitalisation and even patient survival significantly.
Such immediate blood culture results are important in infection prevention and control, and in the early detection of outbreaks (Ombelet et al., 2019). The laboratory should have a system to immediately notify clinicians of significant and urgent results.
Good communication and mutual respect between the clinicians and the laboratory is the foundation stone for building good diagnostic stewardship, as the following activity illustrates.
Before you attempt the activity, note that
Activity 4: CREs and advising clinicians
a.
A highly pathogenic organism is present, so must be treated.
b.
Results are correlated with clinical presentation and a decision is made on treatment.
c.
Bacteraemia is significant, so treatment with meropenem and colistin is necessary.
d.
There is the possibility of poor blood-taking techniques, particularly in neonates.
e.
Infection control measures must be implemented.
The correct answers are b, d and e.
Discussion
Why are these the correct answers?
- Results are correlated with clinical presentation and a decision is made on treatment: We must always treat the patient and use the result in context of clinical assessment and history. This patient is well, so there is no need to treat.
- There is the possibility of poor blood-taking techniques, particularly in neonates: Blood samples obtained from a device can be contaminated, and the volume of blood can also be insufficient. In this case, the patient is well, so it is likely that the CRE was a contaminant. Clinical assessment is the defining factor in the decision to treat: if there is any doubt, a repeat culture should be taken using an optimal technique.
- Infection control measures must be implemented: The presence of a CRE should alert the ICU and the infection control team, because this highly resistant organism is present in the ward and can potentially cause severe infections because it is transmitted horizontally (such as by the hands of ICU staff). Strict infection prevention and control measures should be followed, and regular audits should be in place.
5.2 Treatment does not depend only on laboratory results
It is essential to report results to clinicians appropriately and accurately – but this cannot replace clinical judgement.
The clinician must consider patient-specific factors. For example, if a patient is not responding to the treatment despite the laboratory report indicating that the bacteria causing the infection are susceptible to the recommended antimicrobial agent, there are several possible reasons:
- It could be behavioral: the patient may not be following the instructions and is not taking medications as recommended.
- The patient could be taking a substandard or falsified medicine. Medicines should always be acquired from a reputable source.
- The site of the infection is important: for example, an abscess may require surgical drainage.
- Have you identified and controlled the source of infection? If the source is determined, it may be possible to control the infection by surgical removal. In some cases, the removal of infected intravenous or urinary catheters will control the source and no antibiotic treatment will be required.
- For some infections, such as enteric fever, it takes a few days (72 hours) for the fever to settle after beginning the treatment. Time is required not only for antimicrobial treatment to take effect but for the patient’s immune response to be activated.
5.3 Summary reports on a periodic basis
As well as reporting individual results to clinicians, laboratories should also provide regular reports to the hospital administration and relevant hospital teams and committees. These could include diagnostic stewardship committees, infection control, AMS or drugs and therapeutic committees, or individual departments where AMR is a particular concern such as ICU.
Reporting laboratory results on a regular basis – monthly, for example – has several functions:
- It will provide information on changing patterns of AMR and pathogen incidence in the hospital and locally, which can inform treatment of patients. For example, if resistance to a particular drug is increasing, clinicians will try to avoid using this antimicrobial if alternatives are available. Regular reporting means that the hospital can identify increasing resistance, and formulate and regularly update hospital empirical treatment guidelines.
- It may show new patterns of infection or AMR that require action to prevent an outbreak, such as testing for carriage of a specific AMR organism and increased infection control measures.
- It can be used to monitor the performance of diagnostic stewardship at the facility, and allows a diagnostic stewardship committee to identify performance issues and action areas.
6 Diagnostic stewardship and GLASS
Data collected in the course of clinical diagnostic testing are invaluable in informing institutional treatment guidelines based on local resistance epidemiology. In addition to the test results themselves, demographic information such as age and gender, and how long the patient was in hospital before sampling, is a major source of data that can be used to model the patterns and transmission of pathogens, whatever their AMR pattern.
For the hospital and local healthcare facilities, local or national resistance surveillance data will inform the choice of empirical treatment. However, bacterial resistance patterns are not restricted to geographical regions, but can instead arise and travel rapidly around the world. For this reason, the WHO has implemented the
Figure 5 shows the relationship between laboratory results generated for individual patient care and surveillance data that are used to inform empirical treatment recommendations and AMR control strategies. It illustrates how these can be coordinated at a national level and then fed into GLASS.
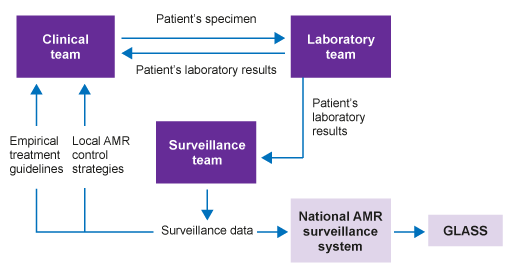
Participation in GLASS requires the use of standard methods for pathogen identification, and AST using CLSI and EUCAST guidelines. It also requires data to be collected at a national level, with both a national co-ordination centre and reference laboratory.
Diagnostic stewardship is a key component for surveillance centres, which supply data to the national system. Guidance for diagnostic stewardship in GLASS is provided by the WHO (2016) and in the associated webinars (Folkhälsomyndigheten Sverige, 2019a, 2019b).
GLASS initially targets only four specimen types: for each one, GLASS provides a list of priority pathogens to be reported. These are reviewed on a regular basis. In addition, GLASS specifies combinations of pathogen and antimicrobials prioritised for surveillance.
It is important to note that GLASS does not require countries to contribute data on every sample, or monitor every priority pathogen. The choice of samples and pathogens can be made in accordance with national priorities and existing capabilities; blood samples are a good starting point for introducing diagnostic stewardship to the standard required.
7 What is needed for diagnostic stewardship?
Organisational aspects of implementing good diagnostic stewardship should begin with a review of existing human, material and financial resources, and an evaluation of the additional requirements.
Cost estimations for each stage of the diagnostic pathway should cover developing and adapting local guidelines and SOPs, and developing and implementing training material, as well as any infrastructure and costs related to efficiently transporting samples, documentation and IT, for example. Table 7 reviews the stages in setting up diagnostic stewardship to the standard expected in GLASS, but can be used for general guidance when GLASS is not the immediate aim.
| Steps | Examples |
|---|---|
| 1. Planning (baseline) | Situation analysis, resources and needs assessment conducted |
| 2. Input (needed resources) | Funding for diagnostic stewardship activities in the surveillance site Local guidelines and SOPs for diagnostic stewardship Trained and capacitated staff on local diagnostic stewardship guidelines Microbiological laboratory facilities with equipment and consumables Communication protocols and facilities |
| 3. Process (activities) | Mobilisation and management of funds Development or adaptation of SOPs Development and implementation of training materials for diagnostic stewardship Implementation of training courses Internal and external quality assurance, regular procurement, and maintenance of equipment and consumables Agreed means and frequency of communication among clinical, laboratory and surveillance staff |
| 4. Output (results) | Sustainable financing and resources available on regular basis Common understanding of protocols for diagnostic stewardship Staff trained and capacitated leading to compliance with local diagnostic stewardship protocols and steps Increase in specimens submitted to the laboratory according to SOPs Good laboratory practices in place resulting in reliable and timely results Patient treatment and surveillance actions are informed in a timely manner |
| 5. Outcome | Patient treatment guided by timely microbiological data resulting in safer and more efficient patient care Accurate and representative AMR surveillance data to inform treatment guidelines and AMR control strategies |
Activity 5: Developing diagnostic stewardship
Watch a six-minute excerpt from Video 1, from 30:03 to 36:00.
If you were to develop a diagnostic stewardship approach appropriate for your healthcare situation, what factors would you need to consider?
Discussion
Any response to this activity will depend on the particular circumstances of your healthcare situation, but you may have thought of:
- costs, in particular the prospective cost
- the availability of sustainable funding
- reimbursement for microbiological diagnostics at your facility
- an appropriate supply chain for consumables and tendering procedures
- available IT systems
- population characteristics – you may need to define the diagnostic approach when there are specific characteristics, which may include a very young or an ageing population, the prevalence of HIV, or the geographical distribution of certain diseases such as malaria, yellow fever or other endemic infections
- AMU in your healthcare facility
- laboratory should establish SOPs for all tests, and each test should be verified or validated before it is introduced
- training and continuous professional development for all staff
- transporting samples between the hospital site and the laboratory
- communication avenues between the hospital site and the laboratory, for example the availability of a clinical microbiologist to visit the wards for discussion with clinicians, and the consistent availability of the laboratory by phone
- awareness among clinical staff of laboratory working hours
- setting up a multidisciplinary team for diagnostic stewardship, in collaboration with infection control and the AMS committee – who should be involved?
This list is not comprehensive; you may have thought of other issues that will affect the implementation of a diagnostic stewardship programme in your healthcare setting.
8 The diagnostic stewardship committee and its role
Managing a patient with an infectious disease requires:
- infection control
- diagnostic stewardship
- antibiotic stewardship.
It is addressed by integrating these three elements, ensuring they are given equal priority. A single committee in charge of all three areas may decide on appropriate sub-structures. Figure 6 shows the close relationship between different stewardship programmes.
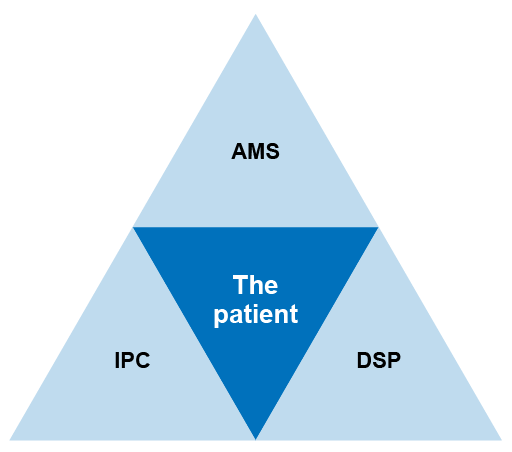
Managing diagnostic stewardship in the hospital or healthcare facility requires a multidisciplinary collaboration between clinical staff, microbiology laboratory staff and surveillance/epidemiological staff. Other staff, such as pharmacists and hospital administration, may also need to be involved.
Although (as mentioned earlier) it might be optimal to combine the different stewardship roles because of the significant overlap, the WHO outlines the role of the diagnostic stewardship committee as follows:
- Development, adoption and implementation of quality management practices including local guidelines and SOPs for specimen selection, collection, transport, laboratory testing and reporting.
- Review and oversight of training needs and activities, including supportive supervision for diagnostic stewardship at the surveillance site.
- Promotion of good diagnostic stewardship at referral sites (particularly if one laboratory in one facility serves several surveillance sites).
- Monitoring of progress of the diagnostic stewardship activities.
- Convening of regular team meetings to (i) present and discuss laboratory results and related issues, (ii) present progress in implementation, (iii) identify and address administrative, technical, operational and logistic issues.
- Establishment of links with the antibiotic stewardship programme, infection and prevention programme, and drug committee.
- Participation in local surveillance data management for reporting and development of local treatment guidelines.
- Key areas of responsibility for each of the different professionals.
Activity 6: Reflecting on how to improve diagnostic stewardship
After studying this module, what are your priorities for improving diagnostic stewardship at your healthcare facility?
Discussion
Your priorities will depend on your role, but options include:
- using posters in wards to illustrate the correct procedure for collecting specimens
- listing information on laboratory working hours and appropriate times for sample delivery in a prominent place in the wards, together with the laboratory phone number
- providing the list of laboratory tests, and brief information on the validation results of new tests that are introduced
- providing short summaries about the purpose of the tests, and their sensitivity and specificity
- introducing regular ward rounds at hospital facilities for the clinical microbiologist, if available, to accompany the clinician
- raising awareness of the local and regional laboratories/hospitals, as well as the local and national governments about the potential for cost savings and improved health outcomes from investing in diagnostic testing and stewardship
- creating guidelines on antimicrobial treatment
- providing periodic information on AMR trends at hospital, ward and patient levels.
This list is by no means comprehensive and the improvements may be simple and specific or more substantial, depending on your role and situation!
9 End-of-module quiz
Well done – you have reached the end of this module and can now do the quiz to test your learning.
This quiz is an opportunity for you to reflect on what you have learned rather than a test, and you can revisit it as many times as you like.
Open the quiz in a new tab or window by holding down ‘Ctrl’ (or ‘Cmd’ on a Mac) when you click on the link.
10 Summary
This module has introduced you to the concept of diagnostic stewardship. At the heart of this is the timely and clear communication between the clinicians and clinical microbiology laboratory personnel, both at the point of taking a correct sample and choosing an appropriate diagnostic test, and in interpreting the results of those diagnostic tests to inform the appropriate treatment for the individual patient.
Diagnostic stewardship encompasses understanding how important it is to submit correct and appropriate specimens in a timely manner, and generating results of the highest quality by the laboratory that are provided to clinicians in a timely and proactive manner.
Depending on the country and local healthcare facility, there will be differences in the initial priorities in setting up diagnostic stewardship; but whatever the baseline level in a particular hospital, patient outcomes can be improved by implementing aspects of diagnostic stewardship, with the additional benefits of providing surveillance data to inform local prescribing practices and reducing the costs of long hospital stays and ineffective treatment.
Diagnostic stewardship is essential for collecting reliable surveillance data locally, nationally and (as capacity increases) internationally for the WHO’s GLASS.
You should now be able to:
- describe the roles in a diagnostic stewardship programme
- understand the principles of taking appropriate clinical samples
- appreciate the range of laboratory techniques available for bacterial isolation, pathogen identification and AST
- understand how microbiology can be appropriately reported to clinicians
- understand factors affecting bacteriology laboratory turnaround times and reporting
- promote good working relationships between laboratories and clinicians for effective diagnostic stewardship
- understand diagnostic stewardship in surveillance at national and international levels
- understand how to introduce a diagnostic stewardship programme.
Now that you have completed this module, consider the following questions:
- What is the single most important lesson that you have taken away from this module?
- How relevant is it to your work?
- Can you suggest ways in which this new knowledge can benefit your practice?
When you have reflected on these, go to your reflective blog and note down your thoughts.
Activity 7: Reflecting on your progress
Do you remember at the beginning of this module you were asked to take a moment to think about these learning outcomes and how confident you felt about your knowledge and skills in these areas?
Now that you have completed this module, take some time to reflect on your progress and use the interactive tool to rate your confidence in these areas using the following scale:
- 5 Very confident
- 4 Confident
- 3 Neither confident nor not confident
- 2 Not very confident
- 1 Not at all confident
Try to use the full range of ratings shown above to rate yourself:
When you have reflected on your answers and your progress on this module, go to your reflective blog and note down your thoughts.
11 Your experience of this module
Now that you have completed this module, take a few moments to reflect on your experience of working through it. Please complete a survey to tell us about your reflections. Your responses will allow us to gauge how useful you have found this module and how effectively you have engaged with the content. We will also use your feedback on this pathway to better inform the design of future online experiences for our learners.
Many thanks for your help.
References
Acknowledgements
This free course was collaboratively written by Olga Perovic and Dawn Harmon, and was reviewed by Priya Khanna, Claire Gordon, Natalie Moyen and Hilary MacQueen.
Except for third party materials and otherwise stated (see terms and conditions), this content is made available under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 Licence.
The material acknowledged below is Proprietary and used under licence (not subject to Creative Commons Licence). Grateful acknowledgement is made to the following sources for permission to reproduce material in this free course:
Images
Module image: KrisCole/iStock/Getty Images Plus.
Figure 1: adapted from Savoldi et al., 2020.
Figures 2, 3 and 6: Olga Perovic.
Figure 4: reprinted from Chabriere et al., 2018, with permission from Elsevier.
Figure 5: WHO, 2016.
Tables
Table 7: WHO, 2016.
Every effort has been made to contact copyright owners. If any have been inadvertently overlooked, the publishers will be pleased to make the necessary arrangements at the first opportunity.