Find out more about The Open University's Health Sciences qualification.
The immune response to viral infection
The immune response to a viral infection is primarily generated by a type of white blood cell called lymphocytes; cells that are mostly localised in ‘lymphoid tissues’ such as the lymph nodes or tonsils. However, the number of lymphocytes that can recognise and react against any individual type of virus is initially very small. This is particularly true for a novel virus such as SARS-CoV-2, which people have never encountered before. In order to produce an effective immune response, the small number of lymphocytes that can recognise a virus must become more abundant. Even though lymphocytes proliferate quickly it still takes several days before there are sufficient cells available to fight back against the infection. During this period the virus may also be spreading rapidly, so there is a race between the virus and the immune system that may determine the final outcome, in terms of recovery.
In the early phase of a viral infection, a variety of immune defences helps to slow viral spread in order to buy time for the lymphocytes to proliferate. One of these defences is mediated by a group of signalling proteins called interferons (Figure 1). When a cell becomes infected by virus, the cell can recognise components of the virus and respond by releasing interferons that diffuse to neighbouring cells and stimulate signalling pathways by binding to receptors on their surface. Consequently, these neighbouring cells start synthesising a large number of anti-viral proteins that put the cells into a state of viral resistance. If a virus ever infects these cells, viral replication is inhibited. By this mechanism, the spread of infection through the tissue is slowed.
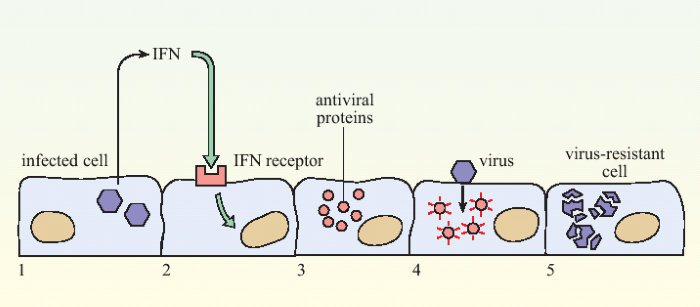 Figure 1. The action of interferon in slowing viral infection.When a cell becomes infected with virus (1) it releases interferons (IFN) which are recognised by receptors on neighbouring cells (2). This cell produces anti-viral proteins (3), so that if it subsequently becomes infected with a virus (4), replication of the virus is inhibited (5).
Figure 1. The action of interferon in slowing viral infection.When a cell becomes infected with virus (1) it releases interferons (IFN) which are recognised by receptors on neighbouring cells (2). This cell produces anti-viral proteins (3), so that if it subsequently becomes infected with a virus (4), replication of the virus is inhibited (5).
Another early-acting immune defence mechanism involves a type of white blood cells called Natural Killer cells (NK cells) – a group of lymphocytes that have the innate ability to recognise virally-infected cells and cause them to die so that the virus can no longer replicate. NK cells recognise alterations that occur in virus-infected cells, although they do not specifically recognise any particular virus infection. These innate immune defences (i.e., interferons and NK cells) are critical in the first few days after infection while the lymphocytes divide and differentiate into cells that are able to mount an effective immune response.
By 3–4 days after infection, there are enough virus-specific lymphocytes available to start mounting an effective immune response against the virus. In order to control the infection, the immune system must both destroy virus-infected cells and prevent the spread of virus between cells. These distinct actions are done by two different sets of lymphocytes.
It is the antibodies that bind to the external spike protein that are most important in limiting virus spread and conferring immunity.Cytotoxic T lymphocytes recognise and destroy infected cells of the body (similar to the NK cells). When a cell becomes infected, the virus hijacks the protein-synthesis machinery of the cell in order to make its own proteins. All cells within the body continually sample the proteins they are producing and place fragments of them on the cell surface for review by passing T lymphocytes. If a T lymphocyte recognises a fragment of a virus on an infected cell it signals to the cell, causing it to commit suicide. If virus replication inside the infected cell is not complete, then this limits production and spread of the virus. B lymphocytes differentiate into antibody-secreting cells, and it is the antibodies that prevent spread of virus between cells.
In essence, the T lymphocytes recognise virus-infected cells, while the antibodies produced by B lymphocytes recognise virus particles in blood and tissue fluids. A molecule that is recognised by a T or B lymphocyte is called an antigen. The proteins that make up a virus are different from those in a host human body. Therefore, T and B lymphocytes can specifically recognise foreign material, such as a viral particle, because of its unique antigens. Let us look now at how this applies to an acute virus infection such as SARS-CoV-2.
In order to infect a cell, a SARS-CoV-2 virus particle must first bind to a specific protein called angiotensin-converting enzyme 2 (ACE2), which is present on the surface of cells in the lungs and other organs such the heart, kidneys and intestines. The SARS-CoV-2 virus attaches to ACE2 via its ‘spike protein’ (Figure 2; for more details see: https://www.ncbi.nlm.nih.gov/pubmed/22816037). Antibodies that bind to the viral spike protein can interfere with its ability to interact with ACE2 and therefore limit the ability of the virus to infect cells . During an infection, antibodies may be produced in response to any of the proteins that constitute the virus, but it is the antibodies that bind to the external spike protein that are most important in limiting virus spread and conferring immunity.
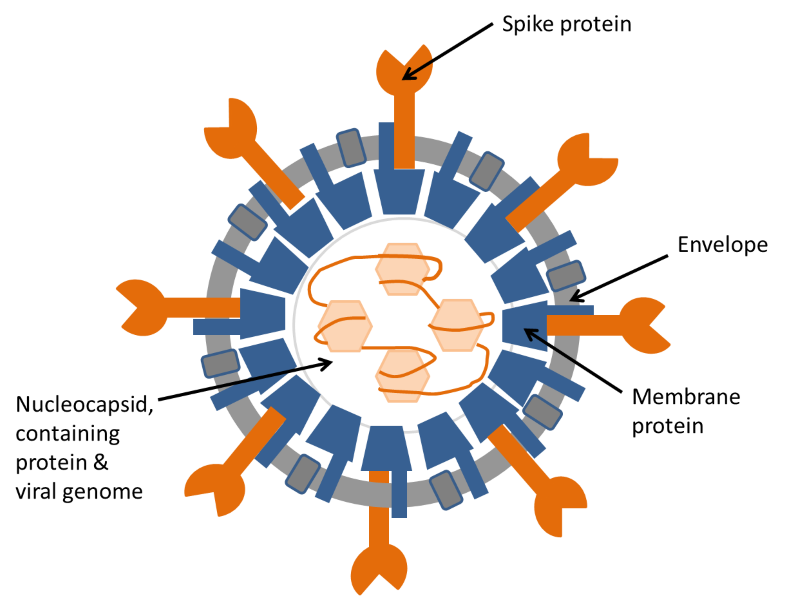 Figure 2. Diagram of coronavirus structure. The illustration shows the structure of a coronavirus in cross-section. The coronavirus genome is a long strand of RNA wound around a protein and assembled inside a shell of membrane protein. This structure is surrounded by an envelope, containing two more proteins, one of which, the spike protein, is responsible for attachment of the virus to the target cell before infection.
Figure 2. Diagram of coronavirus structure. The illustration shows the structure of a coronavirus in cross-section. The coronavirus genome is a long strand of RNA wound around a protein and assembled inside a shell of membrane protein. This structure is surrounded by an envelope, containing two more proteins, one of which, the spike protein, is responsible for attachment of the virus to the target cell before infection.
For more details see: https://www.ncbi.nlm.nih.gov/pubmed/22816037.
The relative importance of the different types of immune defence against viral infection depends on the type of virus. For an acute virus infection such as SARS-CoV-2, immune defences that slow its spread within the body are most important. The main immune defences against acute virus infection are depicted in Figure 3.
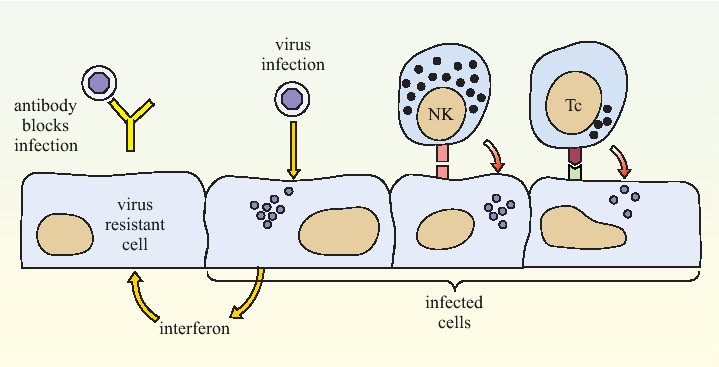 Figure 3. A summary of anti-viral immune defence mechanisms. Antibody binding to the critical antigens on the outside of the virus can prevent it from infecting a host cell. Interferon can signal a cell to adopt a virus-resistant state. Once a cell has become infected, NK cells (NK) and cytotoxic T lymphocytes (Tc) can recognise it and trigger a process of cell death called apoptosis (also known as programmed cell death).
Figure 3. A summary of anti-viral immune defence mechanisms. Antibody binding to the critical antigens on the outside of the virus can prevent it from infecting a host cell. Interferon can signal a cell to adopt a virus-resistant state. Once a cell has become infected, NK cells (NK) and cytotoxic T lymphocytes (Tc) can recognise it and trigger a process of cell death called apoptosis (also known as programmed cell death).
In addition to their role in blocking infection, antibodies also promote removal and destruction of the virus by other cells of the immune system. The production of antibodies that recognise particular viral antigens can persist for many months, which gives extended immunity against a virus, even when all the viral particles are cleared. In the longer term, the production of antibodies that recognise the virus will decline, but the encounter with the virus is not forgotten. The immune reactions that occurred during the viral infection will have caused the population of lymphocytes that recognise that virus to expand. Some of these lymphocytes, referred to as memory cells, retain the ability to react quickly to a repeated infection with the same virus. Hence, long-lasting immunity to a virus infection depends both on antibodies and the memory cells that recognise that virus.
Long-lasting immunity to a virus infection depends both on antibodies and the memory cells that recognise that virus
It is important to note that this continued protection depends on the virus not mutating in such a way that it can evade the immune response. Unfortunately, it is a characteristic of many viruses that they mutate their surface proteins so that the antigens they bear are different and the original antibodies are less effective against them. The mutation rates of RNA-viruses tend to be particularly fast. However, mutation cannot be unlimited since a virus still has to bind to its host cell if it is to survive itself. The internal proteins within a virus are not normally accessible to antibodies, which bind onto the outside of the virus, and they are less susceptible to mutation. Cytotoxic T cells can recognise both the internal and external antigens of the virus provided they are presented on the surface of an infected cell. At this stage (September 2020) we do not know to what extent SARS-CoV-2 can mutate to evade immune responses. Current evidence suggests that it is not a fast-mutating virus by comparison with (for example) HIV. As expected, different proteins of SARS-CoV2 mutate at different rates. In the early months of the pandemic mutations were detected in the spike protein, and two proteins involved in RNA replication, but the rate of mutation of these proteins appears to have slowed. It is thought that these mutations could help the virus evade antibodies, and/or the controls effected by interferons.
An important element of vaccine design is to initiate immune responses against parts of the virus that do not readily mutate. One positive finding is that although many variants of SARS- CoV2 have been identified in different countries, individual antibodies have been found that can bind to most of these variants – such antibodies are said to be ‘cross-reactive’. Ideally a vaccine will induce cross-reactive antibodies which will protect against most or all of the virus variants that might evolve.
You may wonder what makes some individuals more susceptible to severe infections compared to others. Genetic variation in the human population affects all aspects of the immune responses, described above. Hence some individuals will generate faster and more effective immune responses to SARS-CoV-2 than others. However, the immune response is only one component that determines severity of disease. Genetic factors that affect viral entry into the cell and replication are also important. Other factors, such as the initial dose of virus (sometimes called the ‘viral load’), the source of entry and the resilience of the targeted tissues are just as important.
Tests for SARS-CoV-2 infection
There are several methods to determine whether a person has been infected with a virus, which essentially have two objectives:
- Detection of the virus or components of the virus within a person’s cells, blood, secretions, tissue fluids or excretions.
- Detection of an immune response that has been generated by the virus.
For an acute infection like SARS-Cov2, the virus is normally completely cleared from the body around the time that a person recovers from the illness. It is no longer detectable after this time (Figure 4). Conversely, the antibodies that are produced as part of the immune response are first detectable as the patient starts to recover; their production gradually declines over the following months, although they may still be detectable years later. How early the virus can be detected, and how late the antibodies can be detected partly depends on the sensitivity of the assays used.
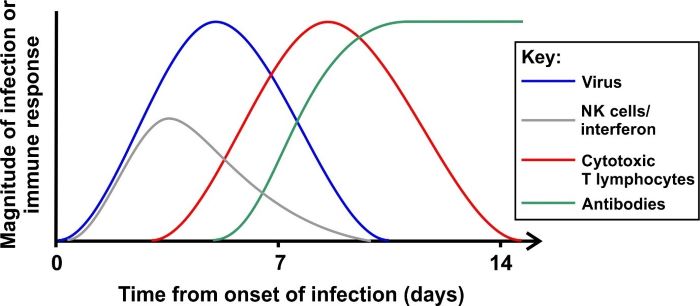 Figure 4. Time course of an acute viral infection and the activity of immune responses The diagram shows the time course of a typical acute infection (although different viruses will vary). A virus is detectable shortly after infection. In the early stages, interferon and NK cells delay viral spread while cytotoxic T cells develop, which can eliminate virus-infected cells. Antibodies that recognised the virus appear as virus levels fall. Antibodies can persist for many months after the infection has waned.
Figure 4. Time course of an acute viral infection and the activity of immune responses The diagram shows the time course of a typical acute infection (although different viruses will vary). A virus is detectable shortly after infection. In the early stages, interferon and NK cells delay viral spread while cytotoxic T cells develop, which can eliminate virus-infected cells. Antibodies that recognised the virus appear as virus levels fall. Antibodies can persist for many months after the infection has waned.
Detection of the virus
It is possible to detect a virus as an infection proceeds, but this is a lengthy procedure that must be done in specialised biological containment laboratories. Routine diagnosis of viral infection looks for viral nucleic acid, which constitutes the genetic material of the virus. It requires careful handling, since the swabs taken from potential patients may have infectious virus on them. The genome of SARS-CoV-2 is a single strand of ribonucleic acid (RNA) a genetic material similar to DNA. The characteristics of any organism is defined by the sequence of bases in its genome. The test for a virus detects just a unique segment of the entire viral genome.
The standard procedure is carried out in three phases:
- Isolation of the viral RNA from swabs taken from the nose or throat.
- Conversion of the RNA sequence into a corresponding DNA sequence (reverse transcription).
- Amplification of the number of copies of the DNA many billions of times to an amount that can be detected – this final step is a qPCR (quantitative Polymerase Chain Reaction).
The first step is a chemical extraction that is often automated. The following two steps are carried out in another instrument called a PCR machine. The specificity of the qPCR (i.e., its ability to amplify only the viral genetic material) depends on the primers that are used. These primers are carefully selected to amplify a segment of the specific viral genome and not DNA from the host or other pathogens that may be on the swab. This procedure takes several hours, and there is no way that this can be reduced without compromising the sensitivity of the test.
A number of new assays have now been developed with the aim of reducing the time taken to test for virus, and/or increasing the number of tests that can be done with the instruments available. One way of reducing the time taken is to combine steps of the assay, or eliminate the need for separate isolation of the viral RNA. Another way of reducing the time is to reduce the number of amplification steps, but this then makes the assay less sensitive.
Another interesting development is the use of ‘Isothermal loop-mediated amplification (LAMP)’. In a normal PCR reaction the samples must be heated, maintained at a high temperature, cooled, and then raised to an intermediate temperature. This cycle must be repeated 30-40 times and that takes the time. In the LAMP technique some very clever molecular biology is used so that the reaction can proceed at one temperature (isothermal amplification). Not only does this obviate the need for the lengthy temperature cycling steps, it can also be done with simpler instruments. The disadvantage is that the reagents required are more complex and the technique is currently less accurate/sensitive than the standard qPCR. Nevertheless being able to reduce the test time to less than 1 hour has major advantages in some settings.
Finally one can consider social aspects of testing. It is much easier for people to provide their own nasal swabs or a saliva sample to test, but such samples are more likely subject to variability and saliva samples generally have less virus to test. This means that higher sensitivity is needed. You can see that there is a trade-off between speed, accuracy and convenience in the different tests. Different parameters are more important for each group of people. For example, a rapid test might be of prime importance in an airport, whereas highest accuracy would be more important in a hospital.
Detection of antibodies
It is important to remember that in any individual there is a time-point when virus can be detected but antibodies have not yet been produced, and a later time period when the virus has been eliminated but the anti-viral antibodies remain.Although there are many elements to the immune response, detection of virus-specific antibodies is the only reasonable way to determine whether a response has taken place. Antibody testing can be applied to large numbers of patients relatively easily. Antibodies appear in the blood as a person recovers from an infection. They may also appear in secretions such as saliva and tears, but detection in secretions is more difficult and may be less reliable. For this reason, the assays detect antibodies in blood (serum; the fluid component of blood). The most commonly used diagnostic test for antibodies is an ELISA (Enzyme Linked ImmunoSorbent Assay). As noted above, antibodies against the spike protein are most relevant to protection against coronavirus infection, but antibodies that recognise other viral proteins (see Figure 2) may be used to establish whether someone has previously been infected with the virus. Each ELISA can quantitate the level of antibodies against one viral protein. For example, to detect antibodies against the spike protein, the spike protein itself would be used as the detection element in the ELISA. An ELISA typically takes ~2 hours to carry out, but this can be abbreviated without greatly compromising the sensitivity of the assay.
There are now many companies trying to develop a lateral flow device (similar to a home pregnancy test). Such tests could give a positive or negative result as to whether an antibody that recognises SARS-CoV-2 is present in the test fluid (e.g., blood obtained by a finger prick) and they can be done in less than 30 minutes. In this case, the test would detect antibody against a SARS-CoV-2 antigen, but unlike ELISA, the test is not quantitative. Tests of this type have been validated for other viruses, but they are normally only used in laboratories. Whether it is possible to develop a test for SARS-CoV-2 of sufficient accuracy is not yet known (September 2020). To read further about this, and to watch a short animation illustrating how a lateral flow device works, read this article from The Conversation .
It is important to remember that in any individual there is a time-point when virus can be detected but antibodies have not yet been produced, and a later time period when the virus has been eliminated but the anti-viral antibodies remain (Figure 4). There is a limited time-window in between when both virus and antibodies can be detected. It is essential that the accuracy of any test is very high, since false-positive results could have serious consequences for the individual; they would think that they were immune to SARS-CoV-2 when they were not.
This article was updated on 25/09/20 to reflect new developments.






















Rate and Review
Rate this article
Review this article
Log into OpenLearn to leave reviews and join in the conversation.
Article reviews